In this tutorial, MatchMaker is used to align protein structures (create a superposition), Match -> Align is used to generate a multiple sequence alignment from the structural superposition, and Morph Conformations is used to morph between related structures.
Sequence alignments are displayed in Multalign Viewer, which is covered in more detail in the Sequences and Structures tutorial, and the morphing trajectory is displayed in MD Movie, which is covered in more detail in the Trajectory and Ensemble Analysis tutorial.
Internet connectivity is required to fetch the structures used in this tutorial: 1tad, 121p, 1r2q, 1j2j, 1puj, 1tnd, 1tag
Protein structures are classified within databases such as SCOP, CATH, and HOMSTRAD. Classifications range from groups of highly similar and closely related proteins to larger, more diverse sets. For analysis and comparison, it is often useful to superimpose related structures. Although it is not always clear whether proteins with the same fold are evolutionarily related (homologous), they should still be superimposable. In general, more closely related proteins are easier to superimpose.
G proteins (guanine nucleotide-binding proteins) are used as examples. G proteins are important in signal transduction. They act as molecular switches, changing conformation and interaction partners depending on whether GTP or GDP is bound. Many diverse structures are known. The two main subsets are the small monomeric G proteins, such as Ras, and the larger heterotrimeric G proteins, which act immediately downstream of G-protein-coupled receptors. The α subunits of heterotrimeric G proteins are homologous to the small G proteins.
Start Chimera by clicking or doubleclicking the Chimera icon
![]() (depending on its location).
Typically, this icon will be present on the desktop.
The Chimera executable can also be run from its
installation location (details...).
(depending on its location).
Typically, this icon will be present on the desktop.
The Chimera executable can also be run from its
installation location (details...).
A splash screen will appear, to be replaced in a few seconds by the main Chimera graphics window or Rapid Access interface (it does not matter which, the following instructions will work with either). If you like, resize the Chimera window by dragging its lower right corner.
Show the Command Line (Tools... General Controls... Command Line). Choose Favorites... Add to Favorites/Toolbar to place some icons on the toolbar. This opens the Preferences, set to Category: Tools. In the On Toolbar column, check the boxes for:
|
|
 |
Fetch a structure from the Protein Data Bank:
Command: open 1tadThe structure contains three copies of the α subunit of transducin, a heterotrimeric G protein. Delete solvent and two of the copies, chains B and C:
Command: del solventMove and scale the structures using the mouse and Side View as desired throughout the tutorial.
Command: del :.b-c
We will superimpose structures of a sample of G proteins, then create a sequence alignment from the superposition.
The α subunit of the heterotrimeric G protein transducin was already opened in the setup. Fetch structures for the monomeric G proteins H-Ras, Rab5a, and ADP-ribosylation factor 1, respectively:
Command: open 121pUse the ribbons preset (which may or may not change the appearance, depending on your preference settings):
Command: open 1r2q
Command: open 1j2j
Menu: Presets... Interactive 1 (ribbons)This preset displays ribbons plus ions, ligands, and nearby sidechains.
| superimposed G proteins |
|---|
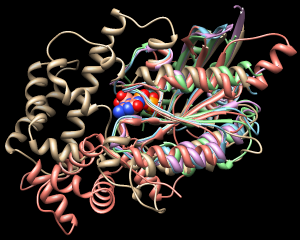 |
The structures need to be superimposed so that they can be compared.
Start MatchMaker
by clicking its icon:
![]()
MatchMaker superimposes structures pairwise by first aligning their sequences and then fitting the α-carbons of residues in the same columns of the sequence alignment. Usually the fit is iterated so that residue pairs aligned in sequence but far apart in space are not used in the final 3D match.
Several parameters control the sequence alignment step:
The number of α-carbon pairs and RMSD in the final iteration of each pairwise fit are reported in the Reply Log (in the menu under Favorites). However, simple visual inspection of the overall structures is often the most useful indicator of success.
Another visual indicator is how well similar ligands superimpose. Show only residues classified as ligand, and label them:
Command: show ligandEach of these structures includes GTP or an analog of GTP in the binding site. However, some other ligands were simply present in the crystallization solution and are not biologically relevant. GOL is glycerol and can be removed:
Command: rlab ligand
Command: del :golTry using different reference structures in MatchMaker (click a line in the Reference structure list, click Apply). With the default alignment parameters, the superposition is similar and basically correct no matter which structure is used as the reference. Detailed examination of the match statistics and guanine nucleotide positions suggests results may be slightly better with 1r2q as the reference.
Command: ~rlab
Next, try a structure that is harder to superimpose, and display its ligand in the sphere representation:
Command: open 1pujBesides lacking sequence similarity, this protein is circularly permuted compared to the others: its N-terminal part structurally matches the C-terminal part of other G proteins and vice versa.
Menu: Presets... Interactive 1 (ribbons)
Command: show ligand
Command: repr sphere ligand & #4
In the MatchMaker dialog, change the Structure to match to only 1puj and try the others in turn as the reference. Again, ligand positions can be used to help gauge the match.
Trials with the default alignment parameters are not successful. When proteins are very distantly related, it may be useful to switch to a lower-number BLOSUM matrix and/or increase the proportion of secondary structure scoring. Usually a range of parameters will give similar results. For example, with 121p as the reference structure, 1puj can be superimposed as shown in the figure with any of BLOSUM 30-75 if secondary structure weighting is raised to 90% and the Smith-Waterman algorithm is used (leaving other settings as defaults). Keep in mind that when proteins are very distantly related, their backbones may diverge even in the best possible superposition.
When all five proteins are superimposed to your satisfaction,
Cancel the MatchMaker dialog.
We will generate a structure-based alignment of the five sequences using
Match
-> Align; start that tool by clicking its icon:
![]()
Match -> Align uses only the distances between α-carbons to create an alignment. Residue types and how the structures were superimposed are not important. All of the A chains should already be chosen in the dialog; the B chain of 1j2j is an unrelated peptide and should not be chosen. Use a cutoff of 5.0 Å, specify Residue aligned in column if within cutoff of [at least one other], and turn on Allow for circular permutation. Click OK to start the calculation.
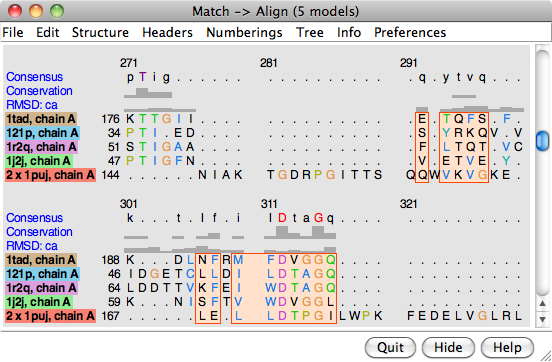 |
It may take a minute or two to create the alignment; progress is reported in the status line. When the calculation is finished, the new alignment will be displayed in Multalign Viewer and can be saved to a file from that tool.
The output multiple sequence alignment (example: 5gees.afa) shows that 1puj was correctly recognized as a circular permutation relative to the others. Match -> Align doubled its sequence to allow C-terminal residues (in the first copy of the sequence) to appear before more N-terminal residues (in the second copy) within the alignment. The columns with residues from all five structures are highlighted as a region in light orange with dark orange outline. Clicking the region will select the corresponding parts of the structures, in effect their common cores. The alignment header named “RMSD: ca” shows the spatial variation per column (α-carbon root-mean-square deviation) as a histogram.
Keep the sequence alignment, but close most of the structures:
Command: sel :/mavPercentConserved=100Some of the conserved residues are Gly (no sidechain). Clear the selection by Ctrl-clicking in an empty area of the graphics window.
Command: disp sel
(To jump to this section right after performing the setup, open the sequence alignment file 4gees.afa included with this tutorial.)
| GTP-binding switch |
|---|
| (1tagA, 1tndA, morph intermediate) |
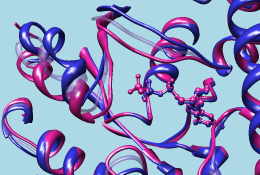

|
Now we will compare 1tad with different structures of the same protein, transducin-α:
Command: open 1tndDelete solvent and chains B-C (extra copies in 1tag):
Command: open 1tag
Command: del solventIf Multalign Viewer (the sequence alignment window) is hidden, bring it to the front by choosing MAV - alignment-name... Raise from near the bottom of the Tools menu.
Command: del :.b-c
Multalign Viewer displays lines of information called headers above the sequences in the alignment. Use the Headers menu to hide Consensus and Conservation and to show RMSD: ca, if not already shown. The sequence name 1tad, chain A has a dashed green line around it, indicating that the sequence is associated with multiple structures. The RMSD header shows the spatial variability of residues associated with each column (α-carbon root-mean-square deviation); currently, it contains high values everywhere because the structures are not all superimposed.
To superimpose the structures using the sequence alignment, choose Structure... Match from the Multalign Viewer menu. One structure (it does not matter which) should be designated as the reference, and all three can be designated as the structures to match. Check the option to Iterate by pruning... using a 2.0-Å cutoff and click OK. The RMSD header is automatically recomputed, showing much lower values.
Superposition of proteins with the same or nearly the same sequence is generally trivial. We used Multalign Viewer since we already had a sequence alignment, but MatchMaker (or its command equivalent) or the command match could have been used instead. These other methods are used and discussed in the Structure Analysis and Comparison tutorial.
Use the ribbons preset (which may or may not change the appearance, depending on your preference settings) and focus on the ligand residues:
Command: preset apply int 1Open the Model Panel and use the S(hown) checkboxes to view the structures individually.
Command: focus ligand
Command: rlab ligand
The 1tad structure (tan) represents the activated form of a G protein; even though it includes GDP, the GDP and ALF (AlF4–) residues together mimic the transition state of GTP hydrolysis. 1tnd (light blue) contains the GTP analog GSP and also represents the activated form. The third structure, 1tag (purplish pink), includes GDP and represents the nonactivated form.
Use the Model Panel checkboxes to show all three structures together. Remove the labels and focus on the overall structures:
Command: ~rlabAlthough the structures are mostly similar, the nonactive conformation (pink) differs from the activated ones (tan and light blue) in specific areas, termed switch regions.
Command: focus
In the sequence alignment window, the three most prominent “humps” in the RMSD header correspond to the known G protein switch regions at approximately residues 173-183, 195-215, and 227-238 of transducin-α. The third switch region is unique to heterotrimeric G proteins; it is an insertion relative to the monomeric G proteins. Placing the cursor over a position in the 1tad sequence lists the associated structure residues near the bottom of the sequence window, and drawing a box around residues in the sequence alignment (click to start, drag to expand) selects the associated parts of the structures.
Close 1tad:
Command: close 0The RMSD histogram looks much the same; now it simply shows the CA-CA distances between the two remaining structures, 1tnd representing the activated form and 1tag representing the nonactivated form.
Finally, morph between the two structures. Morphing involves calculating a series of intermediate structures. In Chimera, the series of structures is treated as a trajectory that can be replayed, saved to a coordinate file, or saved as a movie using MD Movie.
Start the morphing tool:
Command: start Morph ConformationsClick Add... and in the resulting list of models, doubleclick to choose #2, #1, and #2 again, corresponding to a morph trajectory from the nonactivated structure to the activated and back. Close the model list. In the main Morph Conformations dialog, set the Action on Create to hide Conformations, and then click Create.
The progress of the calculation is reported in the status line. When all the intermediate structures have been calculated, the input structures are hidden, the trajectory is opened as model #0, and the MD Movie tool appears.
The trajectory can be played continuously or one step at a time using the buttons on the tool. If the player dialog becomes obscured by other windows, it can be resurrected by choosing MD Movie - trajectory-name... Raise from near the bottom of the Tools menu. If you want to see the original structures again, use the S(hown) checkboxes in the Model Panel.
When you have finished viewing the morph trajectory, choose File... Quit from the menu to exit from Chimera.