Command: mutationscores
The mutationscores command shows the results of deep mutational
scanning on protein structures and as interactive 2D plots.
The Mutation Scores
tool is the corresponding graphical interface.
Deep mutational scanning generally entails making all possible
substitutions (of the 20 standard amino acids) at each position of a protein
and assessing the results with multiple high-throughput assays.
For example, the assays could include growing cells expressing the mutants
in the presence of different drugs and measuring cell viability or fluorescence.
In this page, mutant refers to a variant of the protein
defined by a specific residue type (which could be the same as the wild type)
at a specific position in the sequence, and in which
all of the other positions have the wild-type residue.
The result of a synonymous mutation is the same amino acid
as in the wild-type protein, although the nucleic acid codon could
have been different.
In general, each type of assay yields a score for each mutant.
The scores are read from a mutation scores .csv file
(details...) that can be opened
from the the File menu
or with the open command.
Each file gives rise to a mutation set,
and multiple mutation sets can be open at the same time.
A mutation set can also be fetched
from the AlphaMissense database (score name amiss) or from
UniProt Variants (score names PolyPhen and
SIFT).
Opening a mutation scores file automatically shows the
Mutation Scores tool
for displaying the data as
a scatter plot of the mutants
with two kinds of scores as the axes, or as a
heatmap or histogram
of one kind of score. Each type of plot has controls for changing
which scores are plotted and for which mutation set.
By default, mutation scores are automatically associated with
each structure chain that has exactly the same sequence and residue
numbering (as in the mutation data) when the mutation data or structures
are opened.
However, specific structure chains to associate can be designated with the
chains option of the
open command
at the time of opening the mutation data, or the associations
can be adjusted manually later (after opening the files)
with mutationscores structure.
See also:
Mutation Structure Coloring,
ChimeraX visualization of mutation scores
•
mutationscores heatmap
[ heatmapName ]
[ scores names ]
[ aminoAcids 1-letter-codes ]
[ grouping score | amino acid ]
[ residues residue-spec ]
[ labelEveryResidue true | false ]
[ pixelsPerCell zoom ]
[ palette colormap ]
[ missingValueColor color-spec ]
[ normalizeScores true | false ]
[ subtractFit score-name ]
[ grayMissing true | false ]
[ grayoutColor color-spec ]
[ dragToColor true | false ]
[ dragboxColor color-spec ]
[ dragboxLinewidth pixels ]
[ dragToSelect true | false ]
[ colorResidueOnHover true | false ]
[ hoverColor color ]
[ showOptions true | false ]
[ size width,height ]
[ saveImage pathname | browse ]
[ mutationSet name ]
Create a new heatmap or change the settings of an existing heatmap
with the specified heatmapName.
Default names of heatmap plots are 1, 2, (...).
If an option is not specified for a new heatmap, the default is used,
but for an existing heatmap, the current setting is retained.
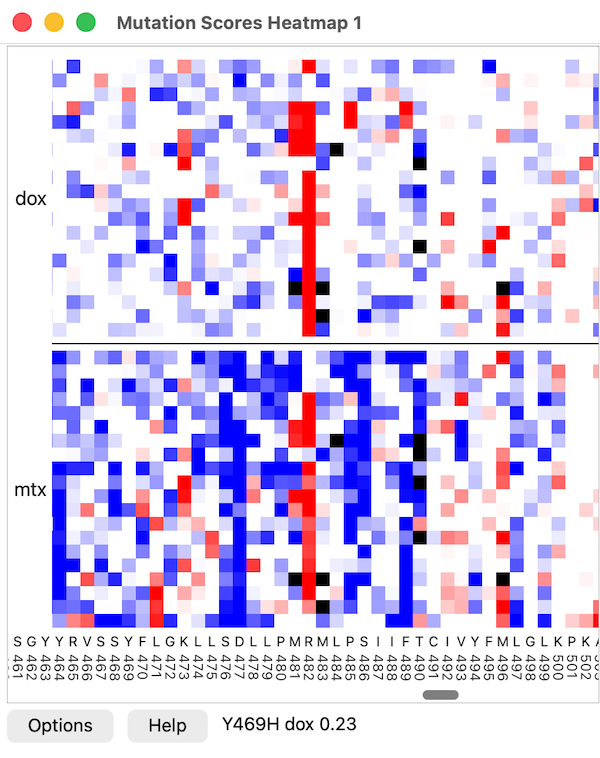
Residue positions along the chain are plotted on the horizontal axis, and
amino acid types along the vertical axis.
Pausing the cursor over the heatmap reports the mutation name,
score name, and value in the status area next to the Help button.
The scores to include are designated as a comma-separated list of
score names.
The aminoAcids option specifies the top-to-bottom ordering of
amino acid types as an uppercase string of amino acid 1-letter codes
(the default HRKDEFWYNQILCSTVMAGP groups chemically similar types);
all 20 standard types may be included, or a subset to limit the plot to
only those types, and the vertical grouping when multiple scores
are plotted can be by score (score name, default) or by
amino acid type. The residues option allows limiting the
plot to only the specified residues,
and the labelEveryResidue option specifies whether to label each
horizontal position instead of at intervals (default false)
Each mutation is represented by a colored square,
with pixelsPerCell specifying the side length of each square in pixels
(default zoom = 2). The coloring palette is given as a
colormap
with value,color pairs (default -2,blue:-1,white:1,white:2,red)
and the color for missing data specified with the missingValueColor
option (default black
).
Data value units are standard deviations above (positive) or below (negative)
the mean if normalizeScores is true (default);
otherwise, the raw values from the data file will be used.
The mean and standard deviation are computed separately for each score
using synonymous mutations only, or if there are
no synonymous mutations, all mutations. Since only a single colormap is used
for the whole heatmap, if raw scores are used, all of the scores should
have comparable ranges.
The subtractFit option allows subtracting
from all scores a linear fit using a specified score.
For instance, for a membrane receptor with scores representing activity in
the presence of different drugs, a linear fit of drug score versus a cell
surface expression score can be subtracted to compensate for different
abundance of the receptor on the cell membrane.
A least-squares fit is used between drug and surface score.
There can be crosstalk between the heatmap and
associated structures.
The grayMissing option (default false)
indicates whether to use the grayoutColor
(default #CCCCCC
)
to modulate the heatmap cell color where
the associated atomic structure is missing residues.
If multiple chains are associated, then mutations are grayed if none of the
chains have the mutated residue. The dragToColor option
indicates whether dragging a box with the mouse on the
heatmap should color the associated structure
residues and atoms (default true, with dragboxColor of
yellow
and dragboxLinewidth of 3 pixels).
By holding the Shift key, multiple boxes can be drawn on
the heatmap, and they can be different colors. Clicking on a mutation
will color a single residue, and clicking outside the heatmap
(such as on the axes) will clear the boxes and revert the residues
to their original colors. If dragToSelect is true (default false),
dragging a box clears the current selection
and selects the associated residues.
If the Shift key is held while dragging, the residues are
added to the selection instead of replacing it.
The colorResidueOnHover option specifies
whether placing the mouse over the heatmap colors the
associated structure residues
(default off, with hoverColor of yellow
).
No mouse button needs to be pressed, and the coloring of the structure
changes immediately when the mouse moves over another mutation.
This facilitates exploring the correspondence between heatmap and
structure positions.
Setting showOptions to true (default false) shows the
options in the heatmap plot window. The size option gives the pixel
dimensions of the visible part of the heatmap within the plot window.
The saveImage option saves the entire heatmap, including axis
scales, to a PNG image file with pathname specified directly
or as the word browse
to specify it interactively in a file browser window.
The mutationSet option allows specifying which dataset to use when
more than one is open. If only one is open, the option is not needed.
The name of a mutation set is derived from the input filename
(for example, abcg2 if read from abcg2.csv).
The names of open sets can be listed in the
Log with
mutationscores list,
and a set can be closed with
mutationscores close.
Currently, a heatmap can only include scores from a single mutation set.
See heatmap details.
•
mutationscores scatterplot
x-score y-score
[ colorSynonymous true | false ]
[ bounds true | false ]
[ correlation true | false ]
[ replace true | false ]
[ mutationSet name ]
Show a scatter plot of mutants for two scores, x-score and
y-score. Only mutants with values for both scores will be plotted.
The colorSynonymous option indicates whether to
color in blue all circles for “mutants” that have the same
residue type as the wild-type protein (default true)
and the bounds option indicates
whether to show dashed lines delimiting mean ± 2 standard deviations
of their values (default true).
The correlation option indicates whether to draw a line
showing the least-squares fit (default false).
The scatter plot will replace a pre-existing scatter plot unless
replace false is given.
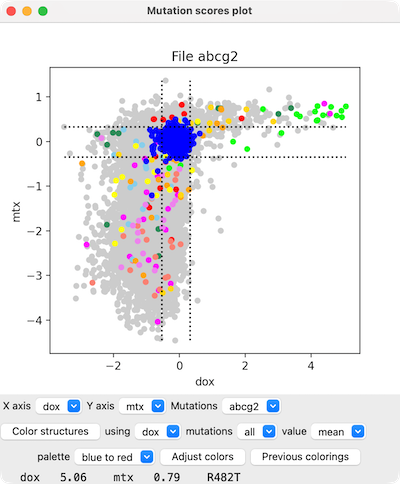
Menus below the plot can be used to change which scores are plotted
on the X- and Y-axes and for which mutation set.
Placing the cursor over a plotted point (circle)
reports the specific mutation and its score values at the bottom of the panel.
See scatterplot details.
•
mutationscores define
[ new-score ]
[ fromScoreName from-score ]
[ aa one-letter-codes ]
[ toAa one-letter-codes ]
[ synonymous true | false ]
[ above value ]
[ below value ]
[ ranges comparison-expression ]
[ combine count | mean | stddev | sum | sum_absolute ]
[ setAttribute true | false ]
[ subtractFit sub-score ]
[ mutationSet name ]
•
mutationscores undefine score
[ mutationSet name ]
A new score can be computed from one or more of the existing scores with
mutationscores define and then used in other
mutationscores analyses just like any of the original scores.
The mutationscores define command without any arguments
will list the existing score names (same as mutationscores
list), and mutationscores undefine can be used to
delete a score.
A new score can also define an interesting subset of the mutations.
The aa and toAa options limit the mutations to specific
“before and after” (before and after mutation) amino acid types.
Specifying synonymous true limits the “mutations”
to those in which the before and after amino acids are the same (wild-type).
The above and below options limit the mutations to those
within a given range of a single type of score specified with
fromScoreName from-score.
This can also be achieved more flexibly, potentially involving more
than one type of score, with the ranges option, which specifies a
boolean expression involving any existing score names along with
"and", "or", "not", ">", ">=", "<", "<=", "!=", and "==" operators and
parentheses, for example, "mtx <= -1.5 and sn38 >= 1.0".
A per-residue score has only a single value per position in the sequence.
Several options create per-residue scores:
synonymous true, or combine, or toAa
with a single amino acid type specified.
The combine option combines the scores for all mutations at a
given position using the specified operation.
With setAttribute true (default), defining a per-residue score
assigns an attribute
with the same name as the score
to the associated structure residues.
An attribute
can be shown with coloring and
used in selection and
and command-line specification.
The subtractFit option performs a linear least-squares fit of the
from-score to the sub-score and subtracts the linear values
from the from-score values.
The typical use of this would be to normalize a score.
For instance, if the from-score is a cell growth measure in response
to a drug and the sub-score is a surface expression score for the
protein, then the subtraction attempts to normalize the drug response to
account for the different amounts of protein reaching the membrane.
A limitation of this simplistic normalization, however, is that the variation
in response might not be a linear function of the quantity of
surface-expressed protein.
•
mutationscores color attribute-name
Reapply a previously defined coloring by mutation score
to the associated structure(s).
A mutation-score attribute must have been
created with mutationscores define
(which assigns an attribute-name same as the score name) and
coloring by that attribute previously applied using the
Render by Attribute tool or the
color byattribute command.
See
Mutation Structure Coloring.
•
mutationscores histogram score
[ scale linear | log ]
[ bins b ]
[ curve true | false ]
[ smoothWidth w ]
[ smoothBins sb ]
[ synonymous true | false ]
[ bounds true | false ]
[ replace true | false ]
[ mutationSet name ]
Show a histogram of mutation scores, with bar-height scaling either linear or
log (default log). The number of bins b in the histogram
defaults to 20. With curve true (default), a smooth curve
is drawn in orange by approximating the histogram with a Gaussian convolution
using smoothWidth w (default 10% of the standard deviation of
the score) and smoothBins sb (default 200).
The synonymous option indicates whether to
show blue histogram bars for the “mutants” that have the same
residue type as the wild-type protein (default true)
and the bounds option indicates
whether to show dashed lines delimiting mean ± 2 standard deviations
of their values (default true).
The histogram will replace a pre-existing histogram unless
replace false is given. The menus below the histogram can be used
to change which score is plotted and for which
mutation set.
See histogram details.
•
mutationscores label
residue-spec
score
[ palette palette ]
[ range low,high | full ]
[ noDataColor color-spec ]
[ height h ]
[ offset x,y,z ]
[ onTop true | false ]
[ mutationSet name ]
Label each of the specified residues in the
associated structure(s)
with a grid of all possible substitutions (the 20 standard amino acids)
color-coded by the values of the score at that position.
The coloring defaults are:
palette redblue
range full
... meaning
across the full range of values (including the values of
residues not being labeled).
A narrower range can be specified as comma-separated values low,high.
The noDataColor is used for residues without mutation scores (default
).
The label height h defaults to 1.5 Å.
The offset x,y,z is the position of the
lower left corner of the grid relative to the residue's α-carbon
along X, Y, and Z in screen coordinates
(default 0,0,3 Å). The onTop setting is whether
to draw the labels on top of other objects regardless of their
relative positions in 3D (default false).
The labels can be removed with
label delete.
•
mutationscores statistics score
[ type all | synonymous ]
[ mutationSet name ]
Compute the mean and standard deviation of a score.
The statistics are reported in the Log.
By default, only the scores of synonymous mutants (proteins with the
same amino acid sequence as the wild type) are included, but type all
can be used to include the scores of missense mutants as well.
•
mutationscores umap score
[ mutationSet name ]
Project the 20-dimensional vector of the scores of the 20 amino acid types
at each sequence position into 2D using UMAP
(Uniform Manifold Approximation and Projection).
In the UMAP plot, each sequence position is shown as a circle labeled with
the residue number and colored by the wild-type residue (20 distinct colors
for the 20 residue types). Only positions at which all 20 residue types
have scores are plotted. This is an experimental feature and thus far,
has not revealed any striking patterns.
•
mutationscores structure
[ list | clear | chain-spec
[ add chain-spec ]
[ remove chain-spec ]
[ allowMismatches true | false ]
[ alignSequences true | false | alignment-ID | sequence ]
[ minimumPercentIdentity percent-ID ]
[ mutationSet name ]
Designate one or more structure chains to be associated with the mutation
scores for further analysis, or list the currently associated chains,
or clear (remove) the current associations.
By default, without any of use of this command,
mutation scores are automatically associated with
each structure chain that has exactly the same sequence and residue
numbering (as in the mutation data) when the mutation data or structures
are opened.
However, this command can be used to manually adjust the associations.
If the add option is used, the newly associated chains will be added
to the currently associated chains. If the add option is not given,
chains that meet the matching criteria replace the currently associated chains.
The remove option allows disassociating specified chains.
Multiple structure chains can be associated per
mutation set.
If the chain does not have exactly the same residue types as the wild type
in the mutation data, however, allowMismatches true is needed to
force the association, and if the residue numbering is different, the
alignment should be specified with the alignSequences option,
with possible values:
- true – align the sequence determined from the mutation
data (using X for any missing residues) with the specified structure chain
using the Needleman-Wunsch algorithm; if multiple structure chains are
specified, attempt aligning with each and discard those with less than
minimumPercentIdentity percent-ID (default 50%)
- false – same as omitting the alignSequences option
- a sequence alignment aleady open in the
Sequence Viewer,
specified by the alignment-ID shown in the title bar of that tool.
The first sequence in the alignment must be the same as the sequence
that goes with the mutation data, and there must also be a sequence
that exactly matches (including numbering) each of the structure chains
to be associated. If multiple structure chains to be associated have the
same sequence, the alignment only needs to contain one copy, and the
alignment may also contain additional sequences not used for association
purposes.
– or –
- the sequence that goes with the mutation data,
to be aligned to the sequence(s) of the specified structure chain(s)
using the Needleman-Wunsch algorithm.
As with sequence align,
a sequence can be given as:
- a UniProt
name or accession number
- the chain-spec
of a structure chain already open in ChimeraX
- plain text of the entire amino acid sequence pasted directly
into the command line
- the sequence-spec of a sequence
in the Sequence Viewer,
in the form:
alignment-ID:sequence-ID
(details...)
The allowMismatches option defaults to true if
alignSequences is used, otherwise false.
Associations can also be specified with the
chains option of the
open command at the time
of opening the mutation scores file.
•
mutationscores list
List the names of the currently available score sets in the
Log.
The names are derived from the input filenames
(for example, abcg2 if read from abcg2.csv).
A set can be closed with mutationscores close.
•
mutationscores close
[ mutation-set-name ]
Close a specified set of mutation scores.
The names of currently open sets can be listed in the
Log
with mutationscores list.
UCSF Resource for Biocomputing, Visualization, and Informatics /
May 2026