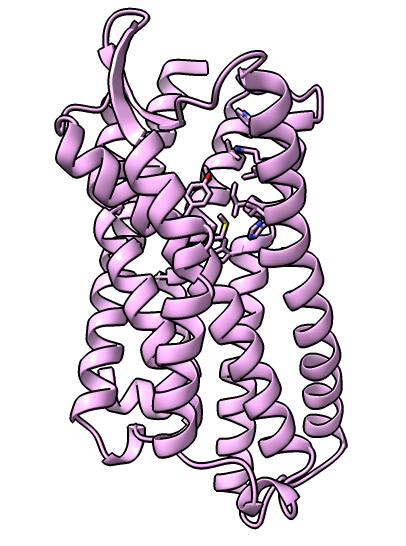
Delta opioid receptor binding of agonist and antagonist ligands
Tom Goddard
January 8, 2020
updated January 5, 2021
This tutorial is for a UCSF Pharm / Med mini-course
DRUG DESIGN: Why is it so hard to design new small molecule drugs?
led by James Fraser.
We will look at how to visualize binding sites including hydrogen bonds and shape complementarity using
ChimeraX software.
The protein we will look at is the delta-opioid receptor, specifically structures described in this November 2019 article
Elucidating the active delta-opioid receptor crystal structure with peptide and small-molecule agonists.
Claff T, Yu J, Blais V, Patel N, Martin C, Wu L, Han GW, Holleran BJ, Van der Poorten O, White KL, Hanson MA, Sarret P, Gendron L, Cherezov V, Katritch V, Ballet S, Liu ZJ, Muller CE, Stevens RC.
Sci Adv. 2019 Nov 27;5(11):eaax9115. doi: 10.1126/sciadv.aax9115. eCollection 2019 Nov.
These are the first delta-opioid agonist structures and the authors suggest they will be useful as a basis
for developing new less addictive pain medicines that target the delta-opioid receptor instead of the
mu-opioid receptor which current pain medicines act on. They say the delta-opioid receptor is also a potential
drug target for anti-anxiety and anti-depressant medications. Therapeutic potential discussed in a 2011
article.
Opioid receptors release a G protein
-
Look mu opioid receptor structure 6ddf to show G protein.
ChimeraX command to fetch 6ddf from PDB:
open 6ddf
- Opioid receptors are GPCR transmembrane proteins.
- They release 3 subunits of a G protein inside the cell when activated.
- Released G protein subunits move along membrane affecting ion channels and cause regulatory signalling that affects gene expression.
|
| 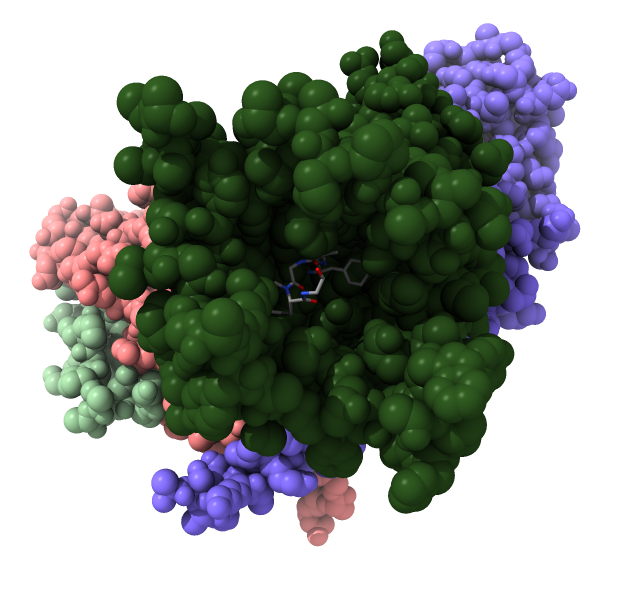
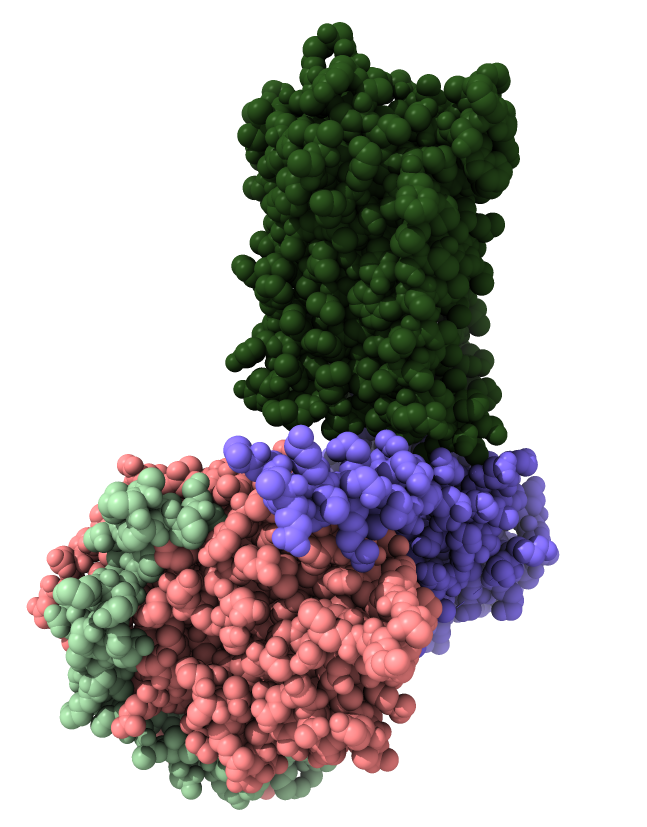
| Opioid receptor is dark green.
G protein subunits are pink, blue and light green.
|
|
Delta opioid receptor with inhibitor bound
|
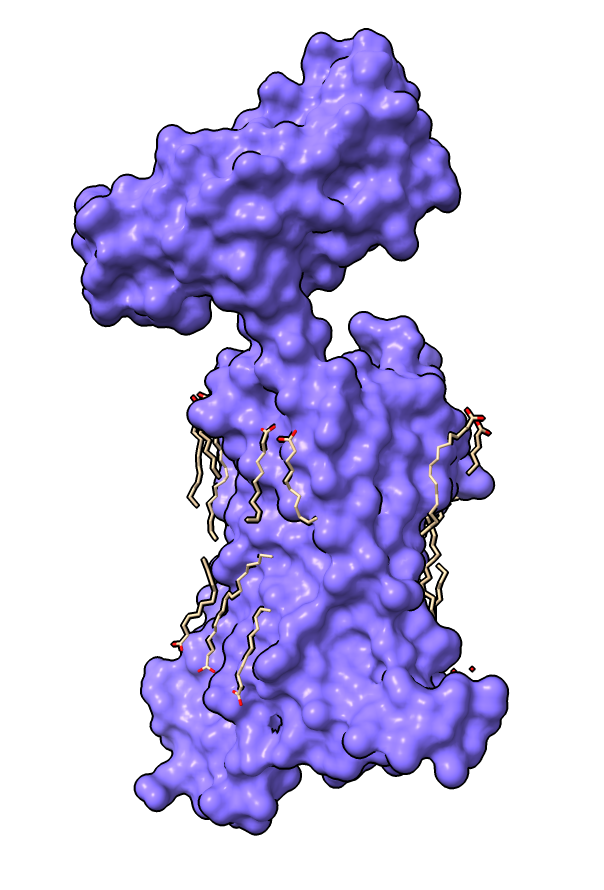
| 
|
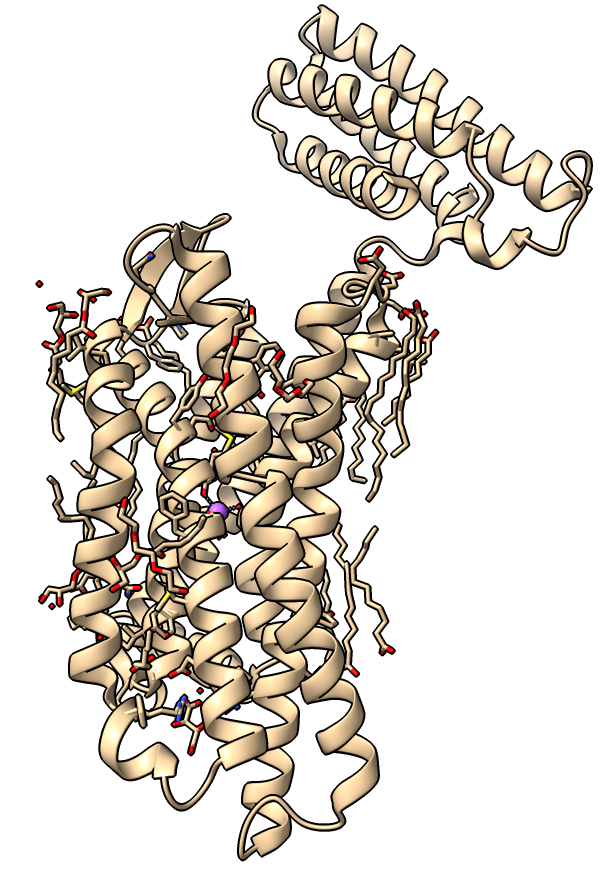
| Delta-opioid receptor with domain at top added for crystallization.
| Surface with adjacent lipids.
|
|
Hydrophobicity coloring
|
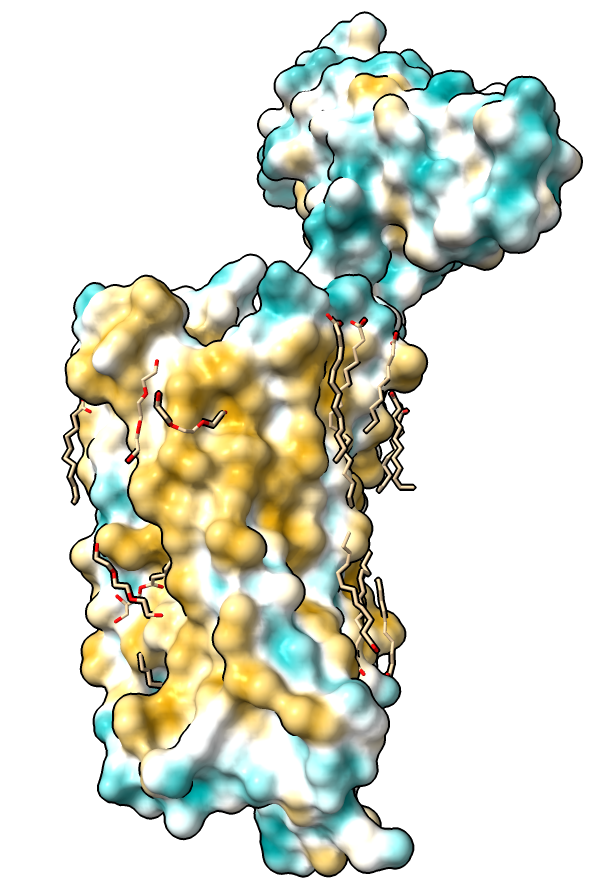
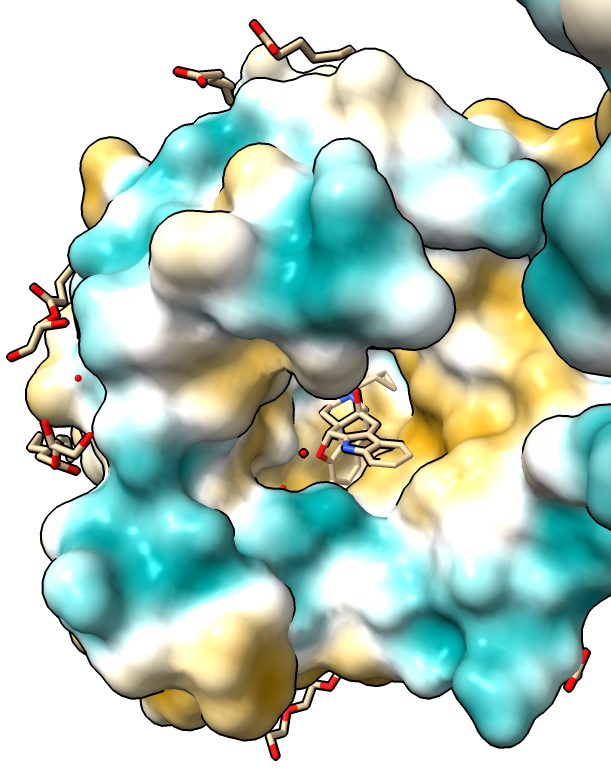
| 
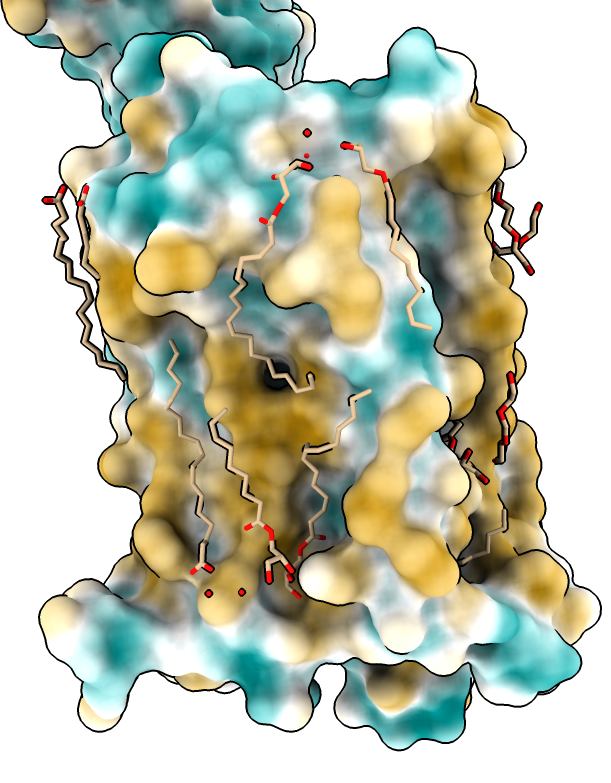
| 
|
| Hydrophobic part yellow is in membrane. Hydrophilic part is blue.
| Naltrindole inhibitor.
| Shadow lighting.
|
|
Hydrogen bonds
- Hide surface
hide surface
- Delete domain added for crystallization, not of interest. Mouse over ends to get residue numbers.
delete :1002-1106
- Show hydrogen bonds (molecule display toobar or command)
hbonds
- Hydrogen bonds only to the antagonist drug.
hbonds :EJ4
- Use clip planes to get better view, menu Tools / General / Side View. Drag vertical clipping lines in side view panel.
|
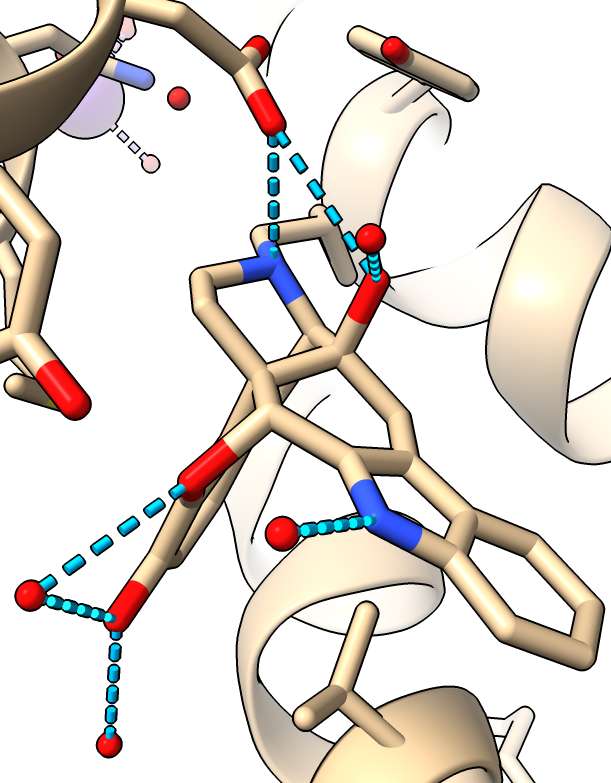
|
| Hydogen bonds blue dashed lines.
|
|
Coloring
- Color drug so it is easier to distinguish from protein.
color :EJ4 orange
- Make oxygens red and nitrogens blue, only carbons stay orange
color byhet
|
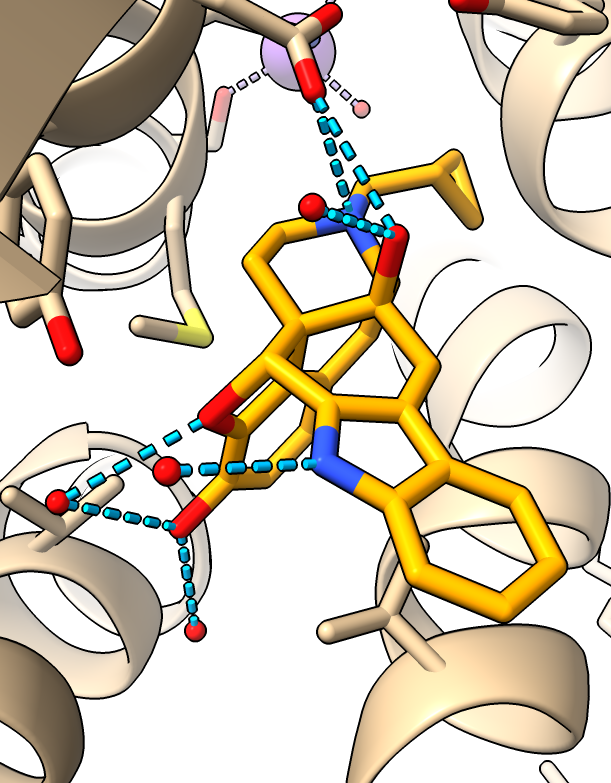
|
| Ligand colored orange.
|
|
Zone
|
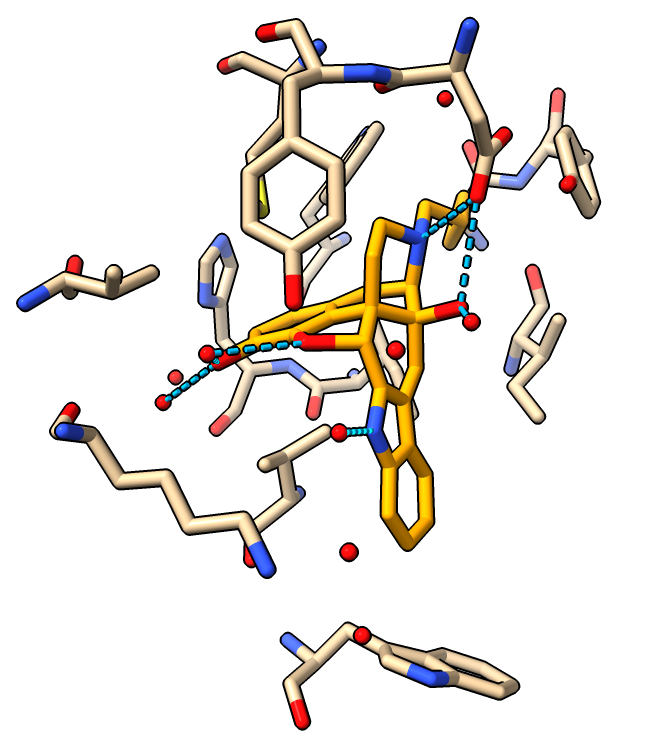
|
| 4 Angstrom zone around ligand
|
|
Add hydrogens
- Hydrogen atoms are usually omitted from experimental structures
because they are too small to resolve.
- To better understand hydrogen bonds add hydrogens.
addh
- Added hydrogens are positioned to optimize hydrogen bonding.
- Most hydrogen bonds with ligand in this structure are to water molecules.
|
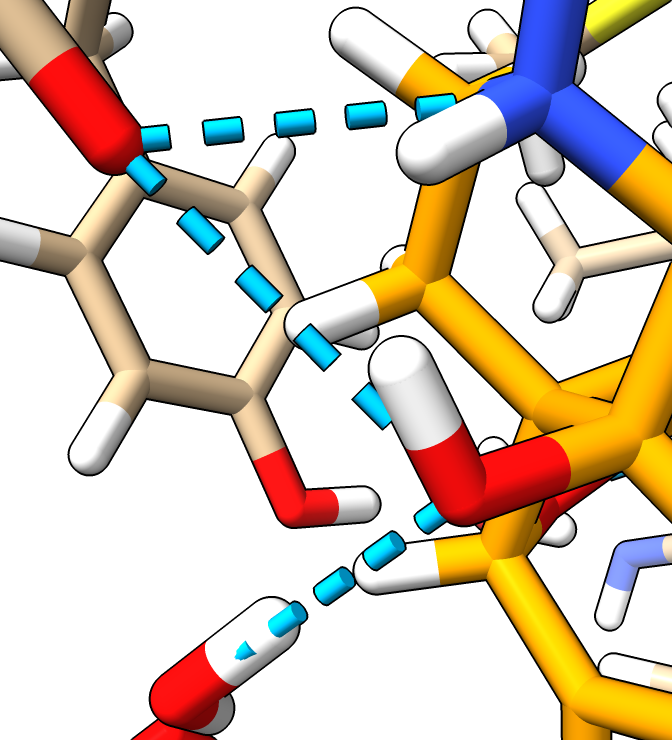
|
| Hydrogens in white.
|
|
X-ray density
|
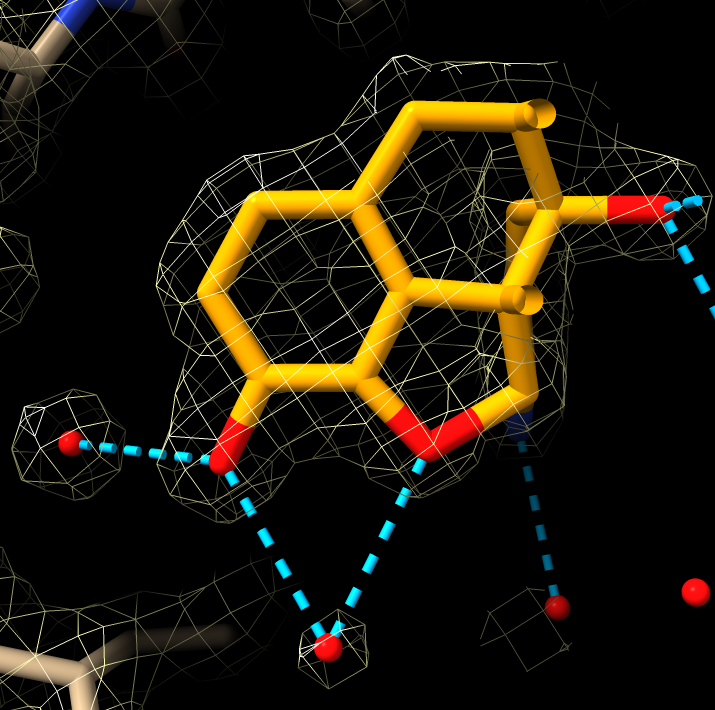
|
| X-ray density.
|
|
Shape complementarity
|
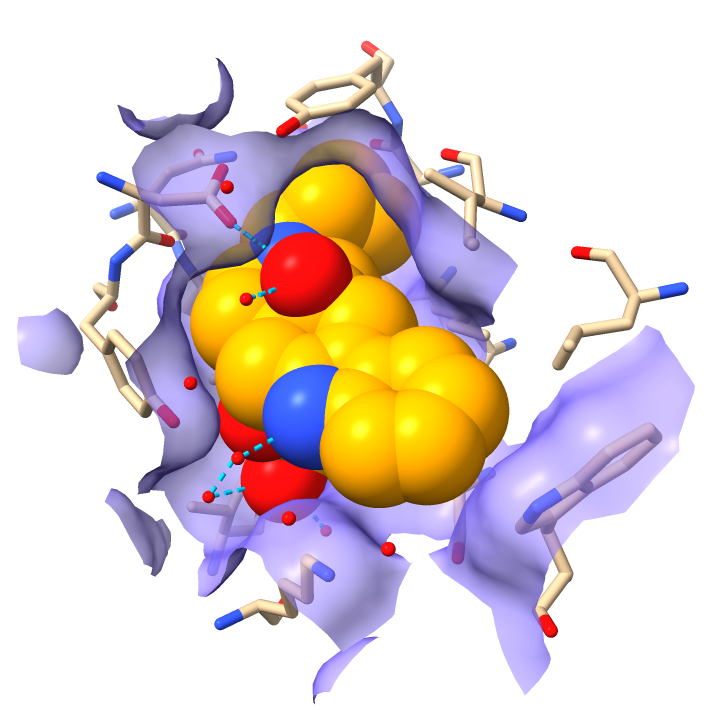
|
| Ligand / receptor complementary shape.
|
|
Activated receptor
- So far looked at inactive receptor with antagonist naltrindole.
- Now lets compare to active receptor with penta-peptide agonist KGCHM07 published Nov 2019.
open 6pt2
hide #1 model # Hide previous model.
view # Center view on new structure.
- There are two copies of the receptor so delete one (chains B and D).
- Also delete the extra crystallization domain.
delete /B,D
delete :999-1106
|
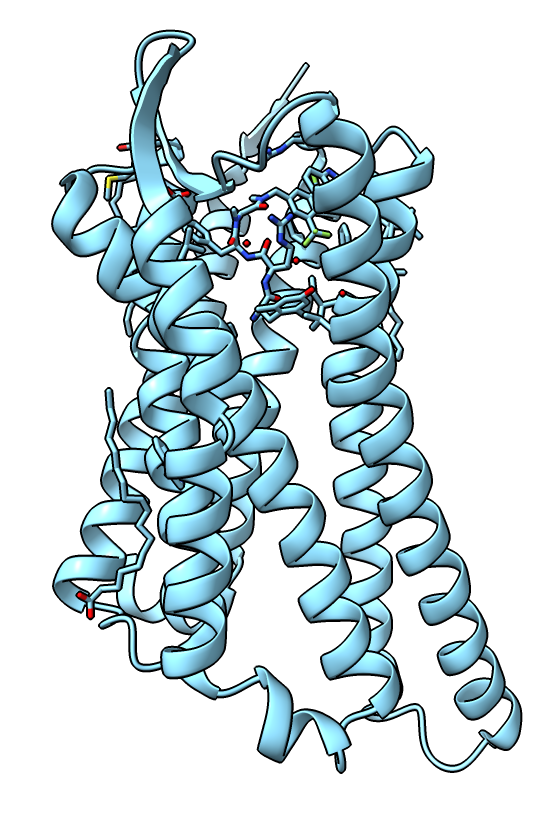
|
| Receptor active state.
|
|
Mutations to stabilize active state
- The authors made 9 amino-acid mutations of the receptor to help stabilize the active state. Show them.
color #2:73,90,95,108,131,143,268,309,323 red
- We see just one, K108 -> D is near the binding site.
- The authors did assays to check that the mutations don't alter the receptor behavior.
|
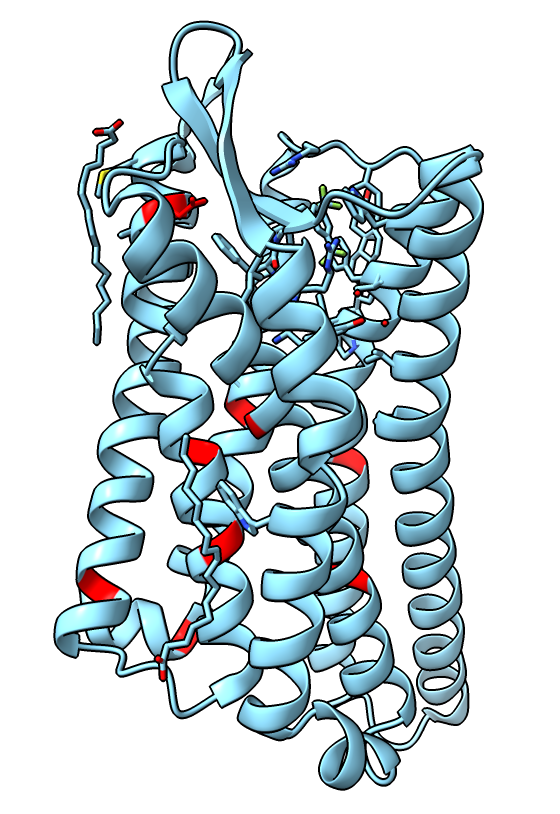
|
| Receptor mutations.
|
|
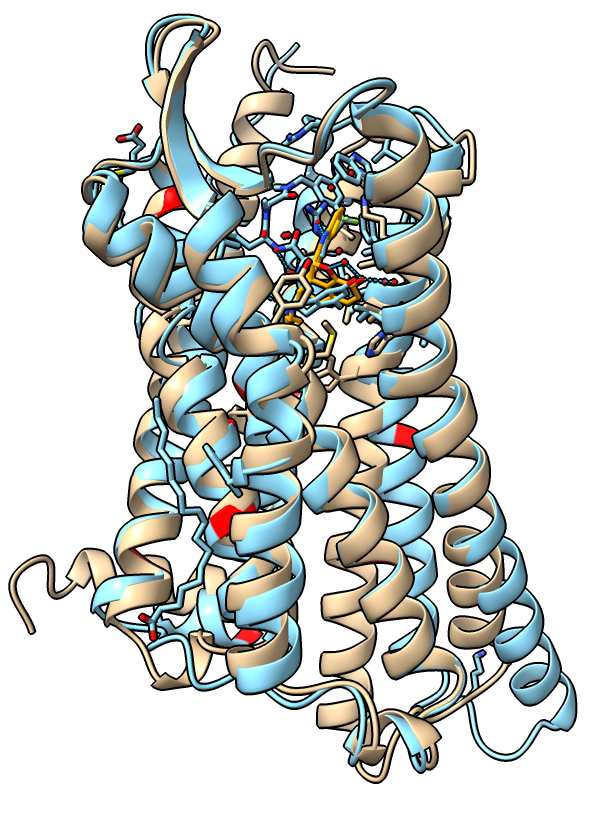
Aligning inactive and active receptor
|
|
| Receptor active (blue) and inactive (tan) states.
|
|
Morphing between inactive and active states
- Another way to see the difference between the two receptor states is to morph.
morph #1,2 same true
- The morph only includes atoms that are present in both structures, so it does
not contain the ligands.
|
|
Critical receptor residue for activation
- The authors find that receptor residue D128 is critical to activation.
Mutating D128 from aspartic acid (D) to asparagine (N) or alanine (A)
make the pentapeptide ligand not activate the wild-type receptor
(measured by an assay where active receptor inhibits cyclic AMP production).
- D128 forms a long range salt-bridge to the peptide ligand.
ChimeraX will show it if the hbond requirements are relaxed.
hide #3 model # Hide morph that does not have ligand
show #2 model # Show 6PT2 with peptide ligand
show :128
color :128 & C yellow # Color residue 128 yellow
color /C & C pink # Color ligand pink
hbonds /C distSlop 0.6
- Can swap aspartic acid to aparagine with swapaa command.
swapaa #2:128 ASN
- To get an asparagine with a better orientation, look at all common rotamers
swapaa interactive #2:128 ASN
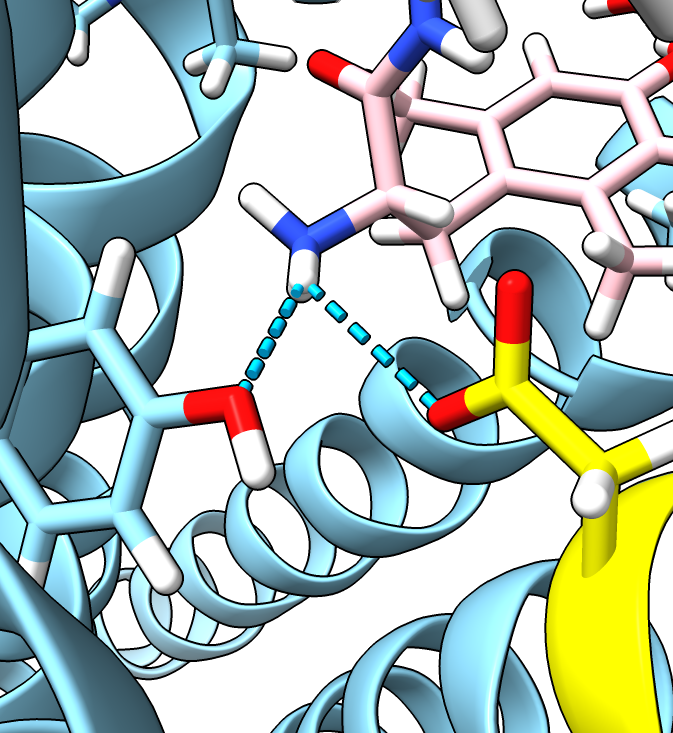
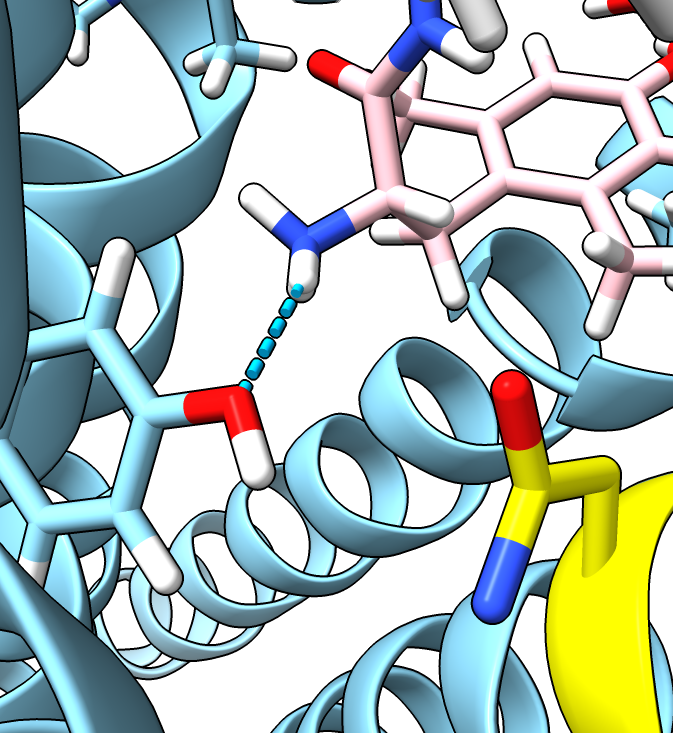
- The second entry in the table with probability 10% is close to the original aspartic acid and
makes the same hydrogen bond. The most probable rotamer 29% does not make the hydrogen bond.
|
| 
|
| Residue D128 yellow, ligand pink.
| D128N mutation (not energy minimized).
|
|