open 6n2y
sequence chain #1/A

Turn off 3-finger mission control on Mac in system preferences Trackpad / More Gestures.
Mouse, left rotate, right translate, scroll to zoom.
log metadata #1
Tom Goddard
January 15, 2020
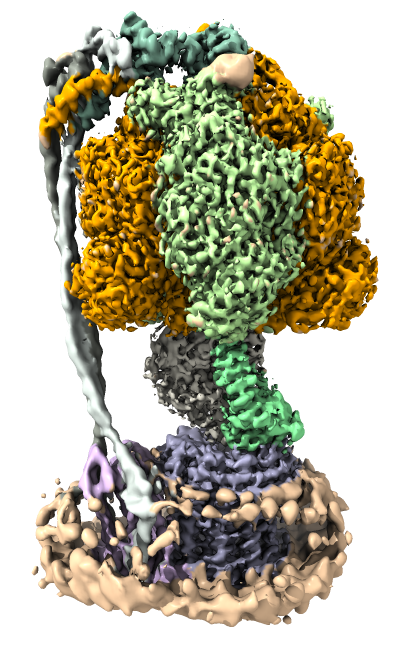
This ChimeraX tutorial will look at how to visualize atomic models and maps of three conformations of a bacterial ATP synthase determined by cryoEM at 3.0 and 3.2 Anstrom resolution. We will create views of the data similar to those found in
Structure of a bacterial ATP synthase.
Guo H, Suzuki T, Rubinstein JL.
Elife. 2019 Feb 6;8. pii: e43128. doi: 10.7554/eLife.43128.
Atomic models PDB 6n2y, 6n2z, 6n30. EMDB maps 9333, 9334, 9335. ChimeraX can fetch these data files directly from the PDB and EMDB or you can download the files for offline use.
Command links in this page will execute in ChimeraX if this web page is displayed within ChimeraX. To show this tutorial in ChimeraX use menu entry Help / Documentation Index, choose Tutorials, and select the January 2020 cryoEM tutorial.
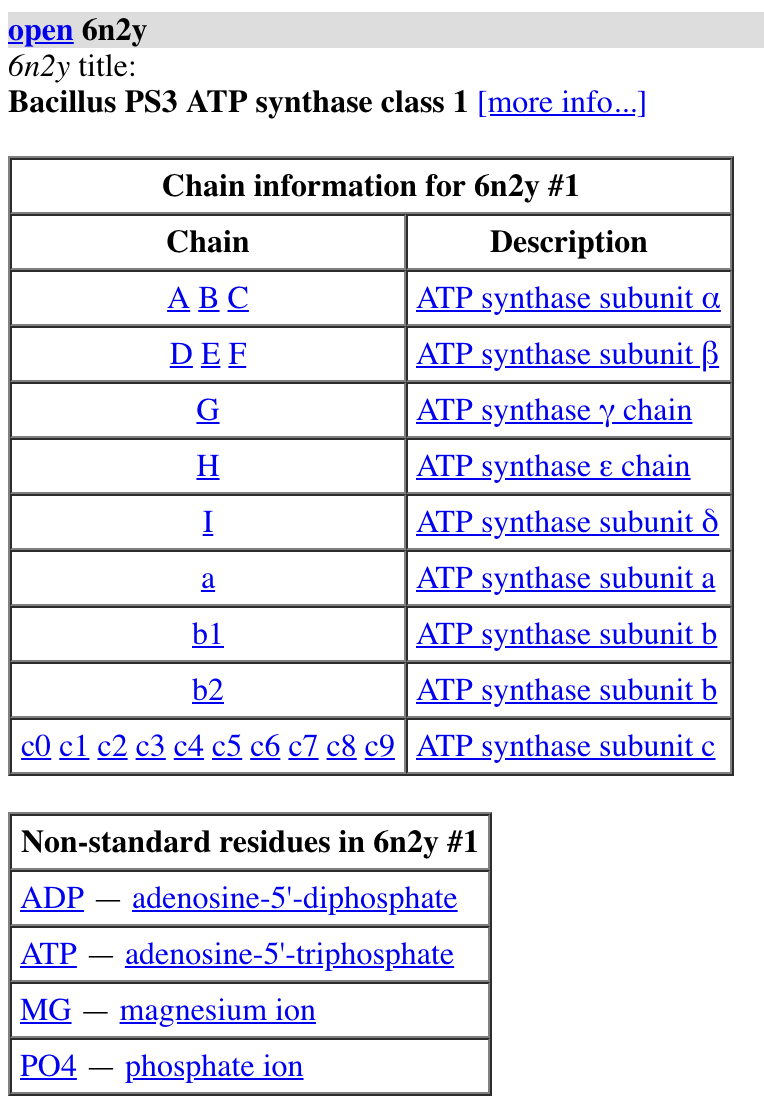
open 6n2y
sequence chain #1/A

log metadata #1
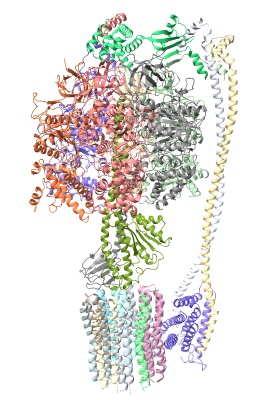
hide atoms show cartoons
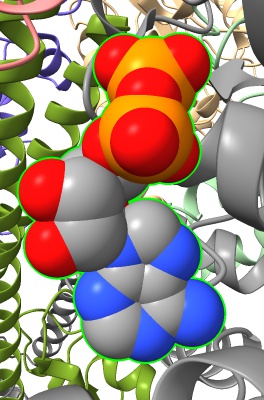
select :ADP show sel atoms style sel sphere
select /H color sel red
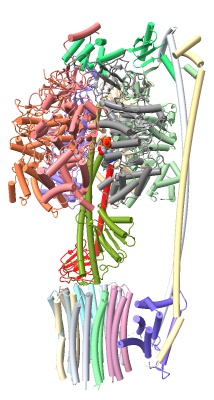
preset cylinders save ~/Desktop/image1.png preset ribbons
open 6n2z open 6n30
morph #1,2
hide cartoon show atoms light simple coordset #4 1,21
close #4
morph #1,2,3 wrap true same true
movie record ; coordset #4 1,61 ; wait 61 ; movie encode
close #2-4 show #1 model
open 9333 from emdb
show atoms style stick
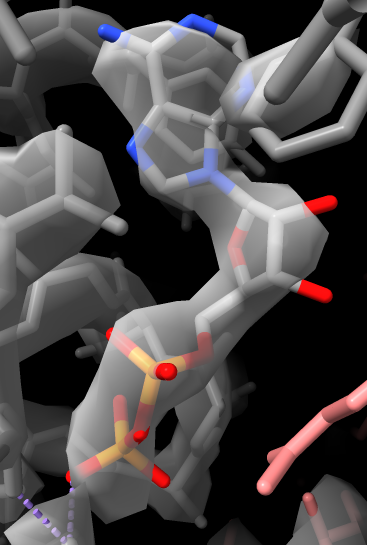
volume #2 step 1
transparency 50
view :ADP
volume #2 level 3
volume #2 style mesh
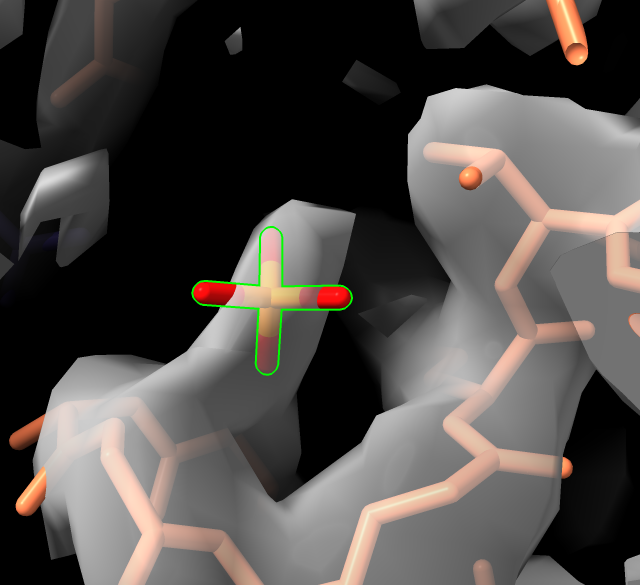
view :PO4
volume #2 level 1
view
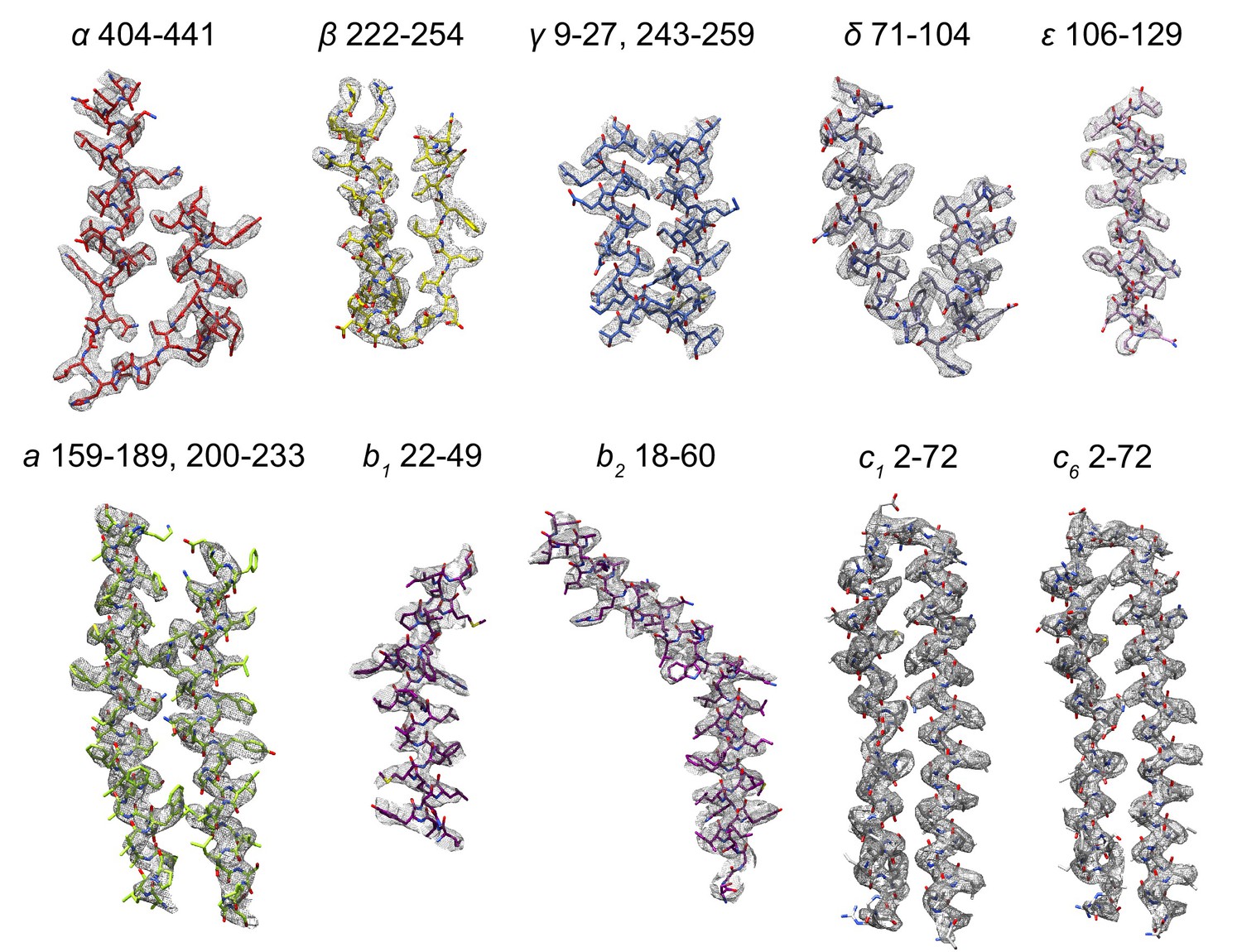

surface zone #2 near #1/A:404-441
hide #1 show #1/A:404-441 view
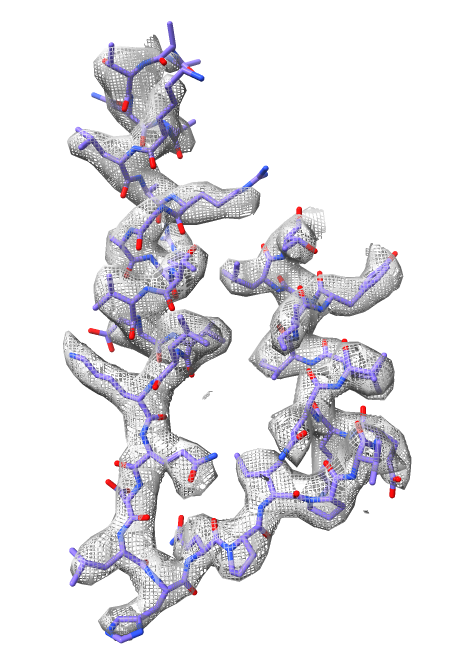
color byhetero
volume #2 style mesh
volume #2 subdivide true volume #2 subdivide true subdivisionlevels 2
volume #2 level 1.5
set bg white
vol #2 subdivide false surface unzone #2
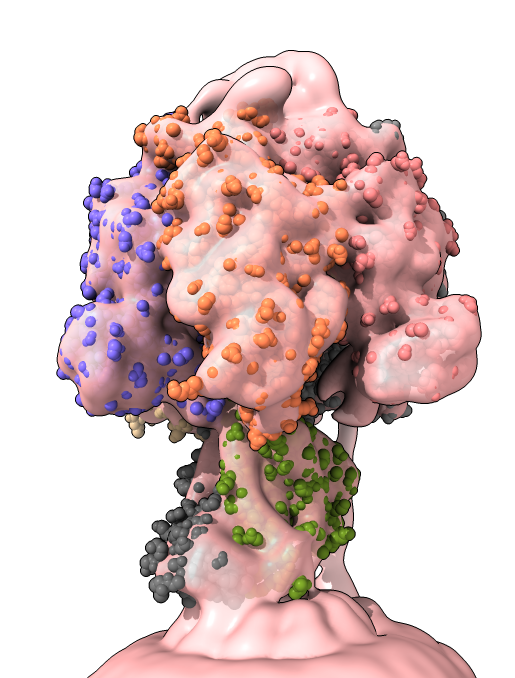
open 4xd7
select #3
mousemode right "translate selected models" mousemode right "rotate selected models" view matrix models #3,0.15538,-0.72458,0.67144,118.99,0.67504,-0.41837,-0.60769,155.34,0.72124,0.54767,0.42412,175.3
fit #3 in #2
volume gaussian #2 sdev 3 fit #3 in #4
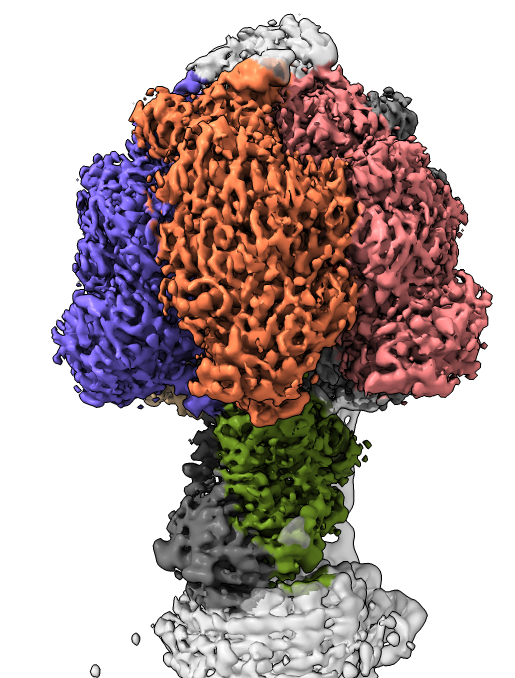
close #4 show #2 model transparency 0 color zone #2 near #3 dist 8 hide #3 model
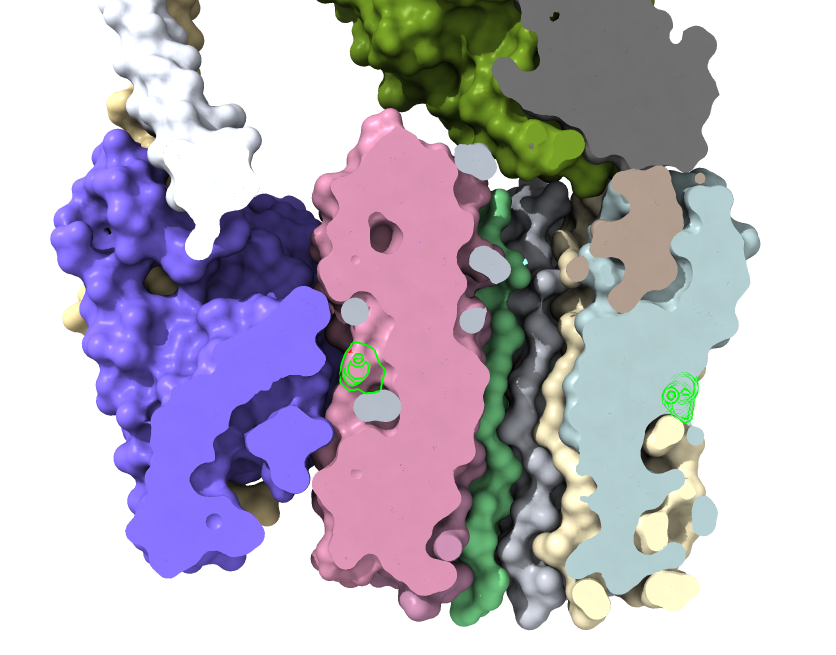
| 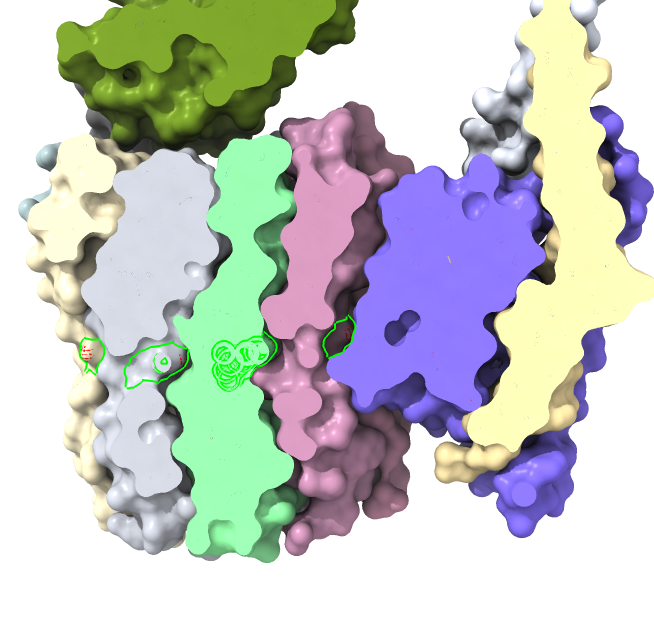
|
| Proton intake channel. | Proton outlet channel. |
close #2,3
show #1 atoms,model select /c*:56 style sel sphere
surface #1
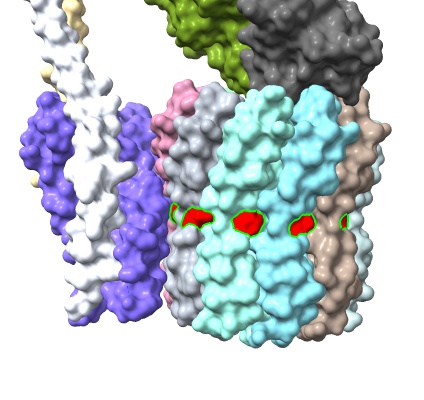
color sel red
mousemode right clip
light simple
clip position 163,163,194 axis -.99,-.08,-.13 coord #1
clip back 0 position 147,162,192 axis .99,.08,.13 coord #1
light soft
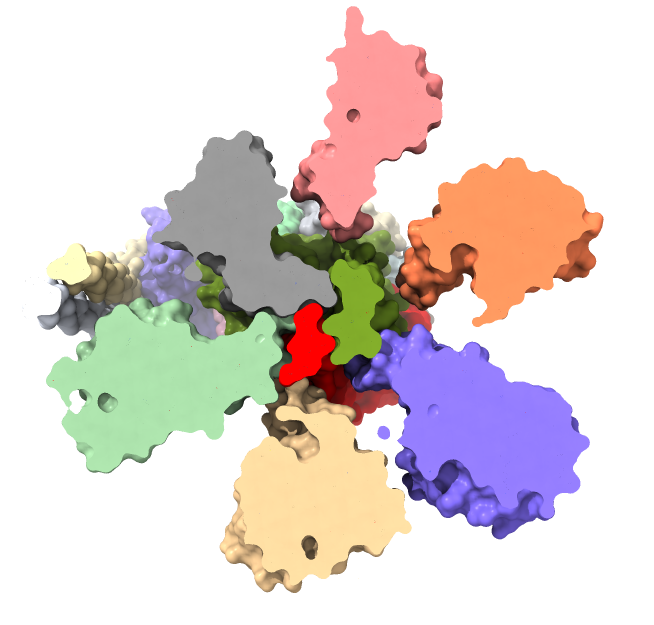
clip off light simple
color /H red
clip axis 0.03,0.13,-0.99 position 164,161,206 coord #1