ChimeraX VR encapsulin ferritin demo
Tom Goddard
Feb 17, 2022
for NIH Virtual and Augmented Reality Scientific Interest Group
Show using ChimeraX virtual reality to look at molecular structures and electron microscopy maps.
Look at encapsulin from Haliangium ochraceum which forms an icosahedral shell around encapsulated
ferritins that convert Fe(II) to the less toxic Fe(III) oxidation state and store it.
There is a video of this demonstration.
Pore dynamics and asymmetric cargo loading in an encapsulin nanocompartment.
Ross J, McIver Z, Lambert T, Piergentili C, Bird JE, Gallagher KJ, Cruickshank FL, James P, Zarazúa-Arvizu E, Horsfall LE, Waldron KJ, Wilson MD, Mackay CL, Baslé A, Clarke DJ, Marles-Wright J.
Sci Adv. 2022 Jan 28;8(4):eabj4461. doi: 10.1126/sciadv.abj4461. Epub 2022 Jan 26. PMID: 35080974; PMCID: PMC8791618.
Data sets EMDB 12873, PDB 7oe2, PDB 5n5f.
Demonstration Steps
Do these steps in VR while screensharing RealSense D435 depth-sensing camera view
showing person and room and models.
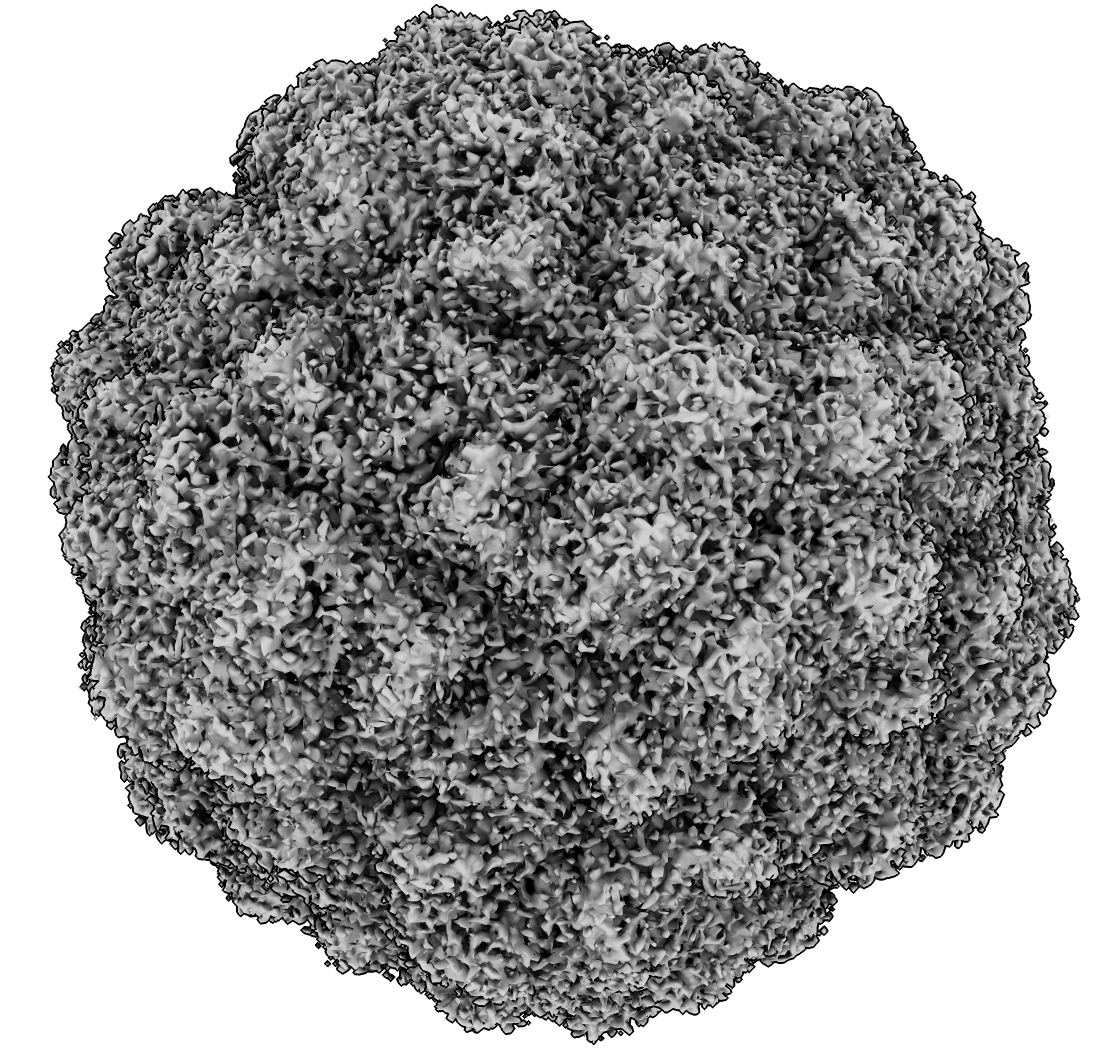
Asymmetric CryoEM Map of Encapsulin Nano-particle
- Start VR with no models open. Make sure all needed models are in file history.
- Make sure RealSense camera Vive Tracker is turned on, then turn on depth camera (command "real on").
- Open EMDB 12873 using file history thumbnail in VR.
- Note this work was published a few weeks ago in Science Advances.
- Explain this is a nano-compartment made of encapsulin protein.
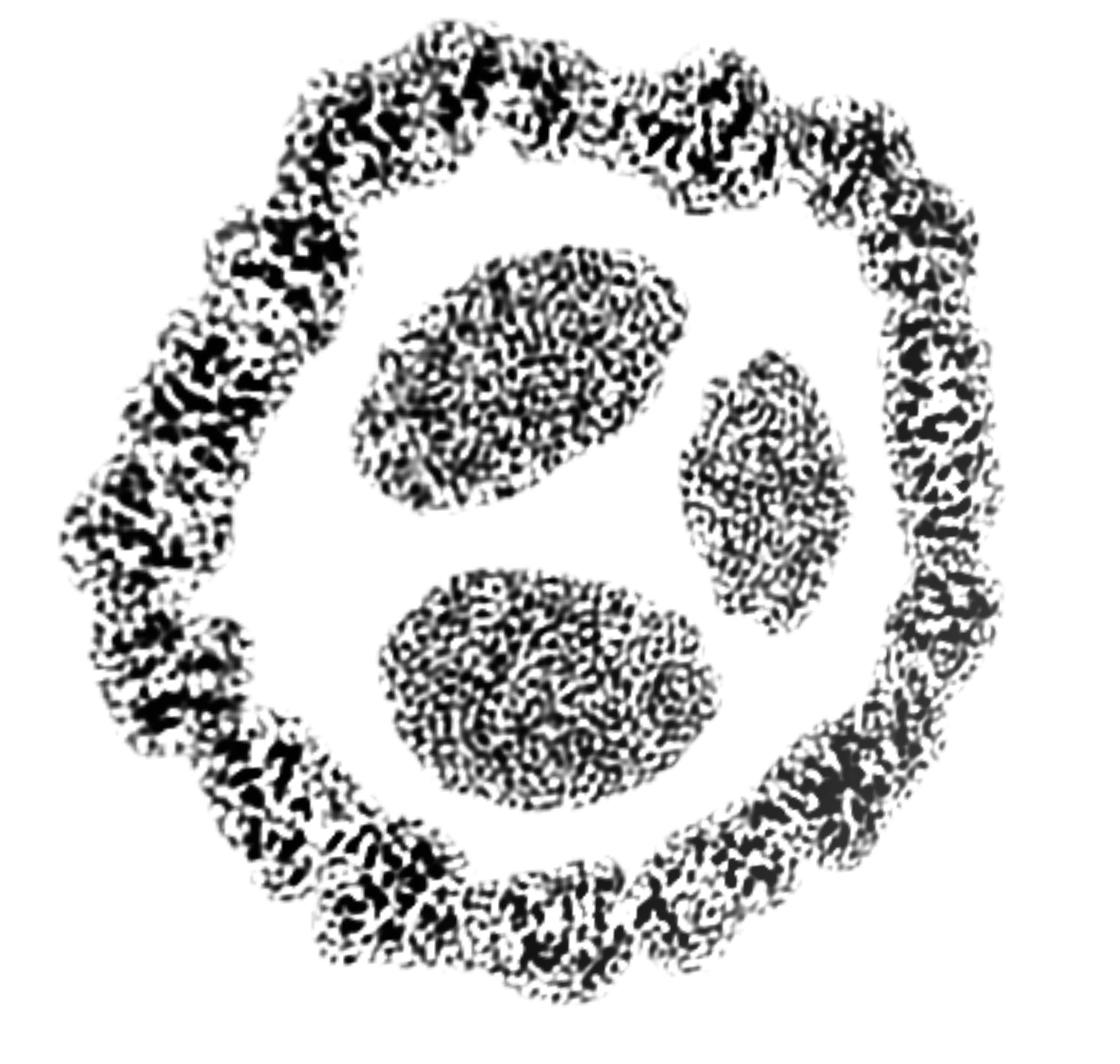

Seeing Inside the Encapsulin Icosahedron
- It contains 4 ferritin complexes that convert Fe(II) to Fe(III) and sequester it.
- To see inside show planes using Map toolbar.
- Use Move Planes mouse mode to slice through it.
- Return to surface display with Map toolbar.
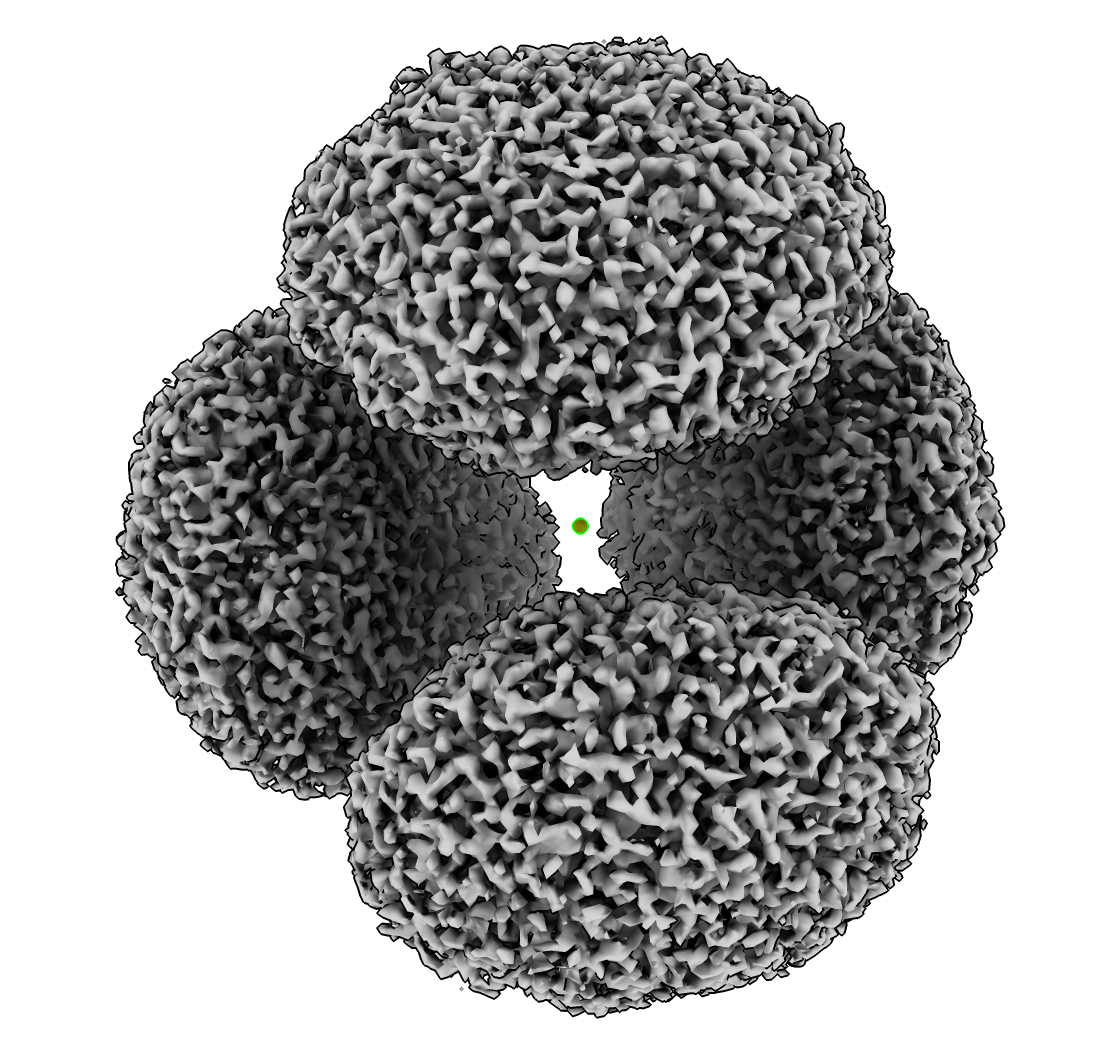
Clip Away Encapsulin Shell
- Show how to remove encapsulin shell to show 4 ferritin complexes.
- Place marker at center with Markers toolbar marking center of connected surface.
- Use menu Tools / Volume Data / Surface Zone to show surface near marker.
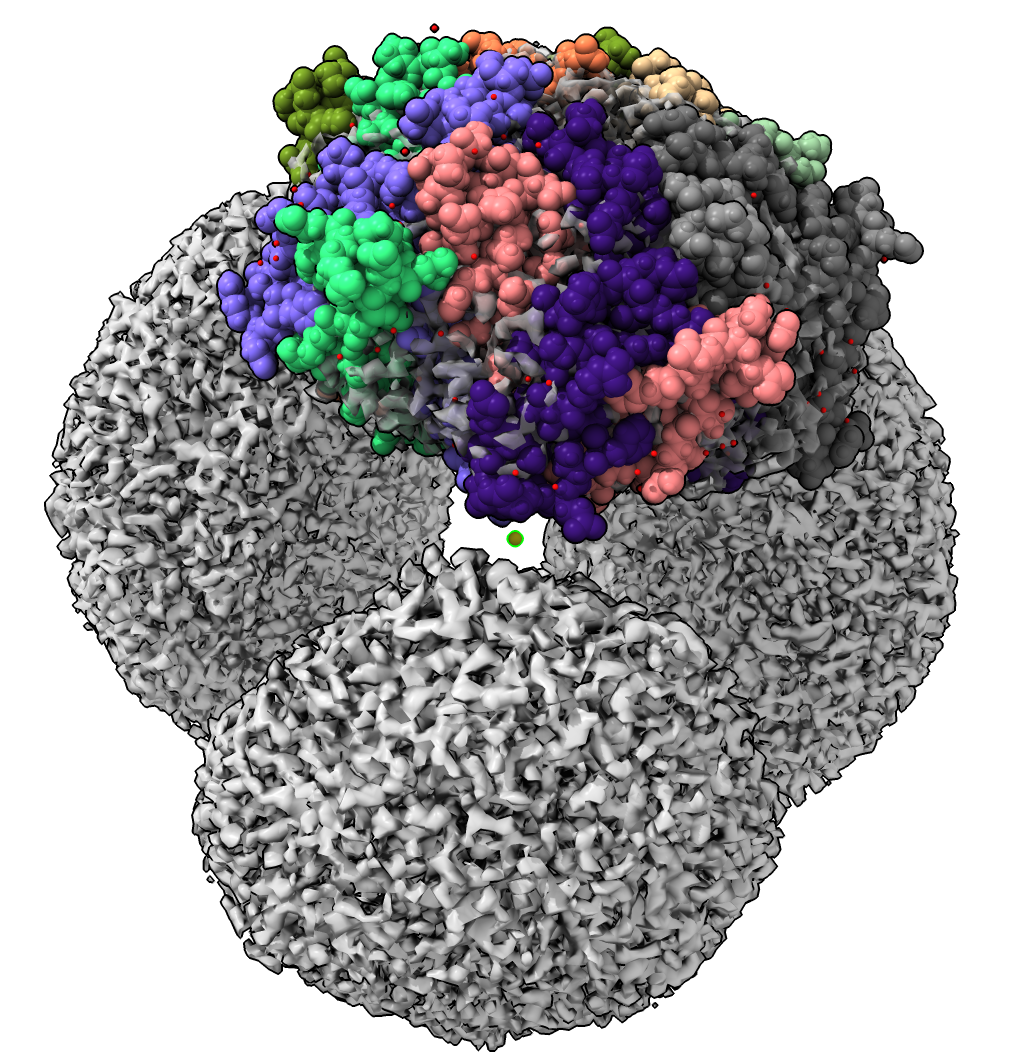
Ferritin Complex Atomic Model
- How do the ferritin complexes end up inside the shell?
- Look at atomic models to understand this.
- Open PDB 5n5f from recent file thumbnails.
- Use drag mouse mode to position it in map.
- Show optimizing position with Map toolbar. Explain that smoothing map needed to get best fit but we will skip that to save time.
- The ferritin complex is a decamer, 10 copies of one protein.
- A localization signal of 9 amino acids is near the C-terminus.
- The localization signal binds to the inside of the encapsulin shell.
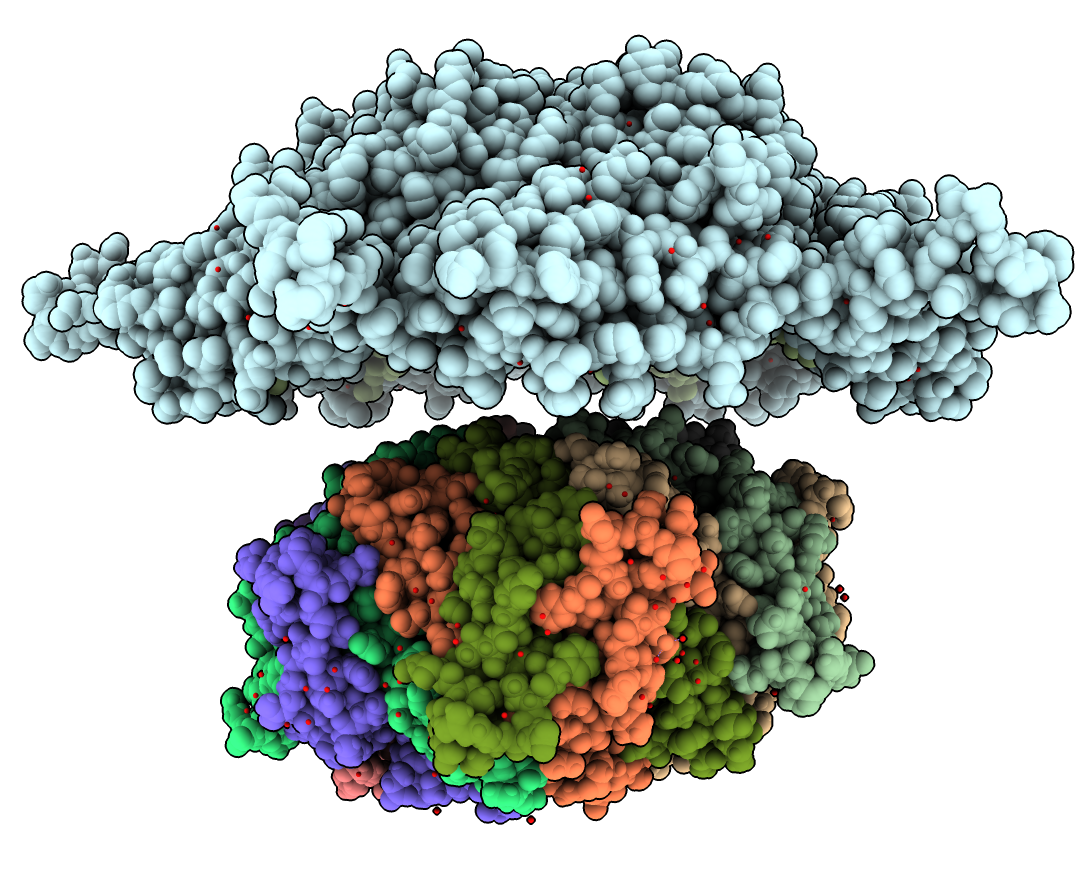
Ecapsulin Shell Atomic Model
- Open PDB 7oe2, a pentamer of the encasulin shell.
- Show the underside with the localization fragment 9 amino acid chain.
- Hand dock pentamer over the ferritin complex after showing the full map with Map toolbar.
- Hide the map with the Model Panel, or Map toolbar.
- Explain that the ferritin experimental structure is missing the localization signal and linker because they are disordered, so not seen in the 5n5f X-ray map.
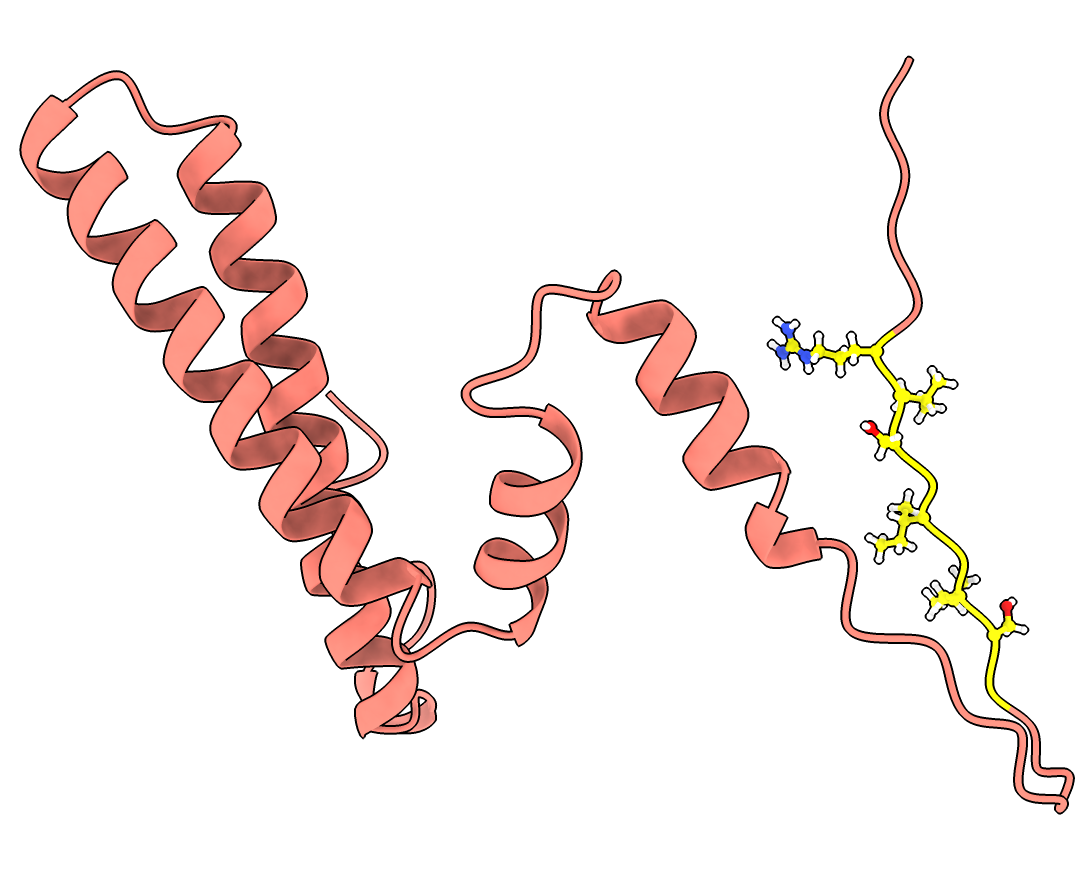
Use AlphaFold Model to get Localization Signal Peptide
- Explain that I ran AlphaFold on a monomer of the ferritin complex to get a model with the localization signal.
- Amazingly AlphaFold predicted the exact conformation even know it forms an intertwined dimer. Magic!
- Open the AlphaFold model and hand align it to a 5n5f chain, showing both as ribbon.
Connect Ferritin to Shell
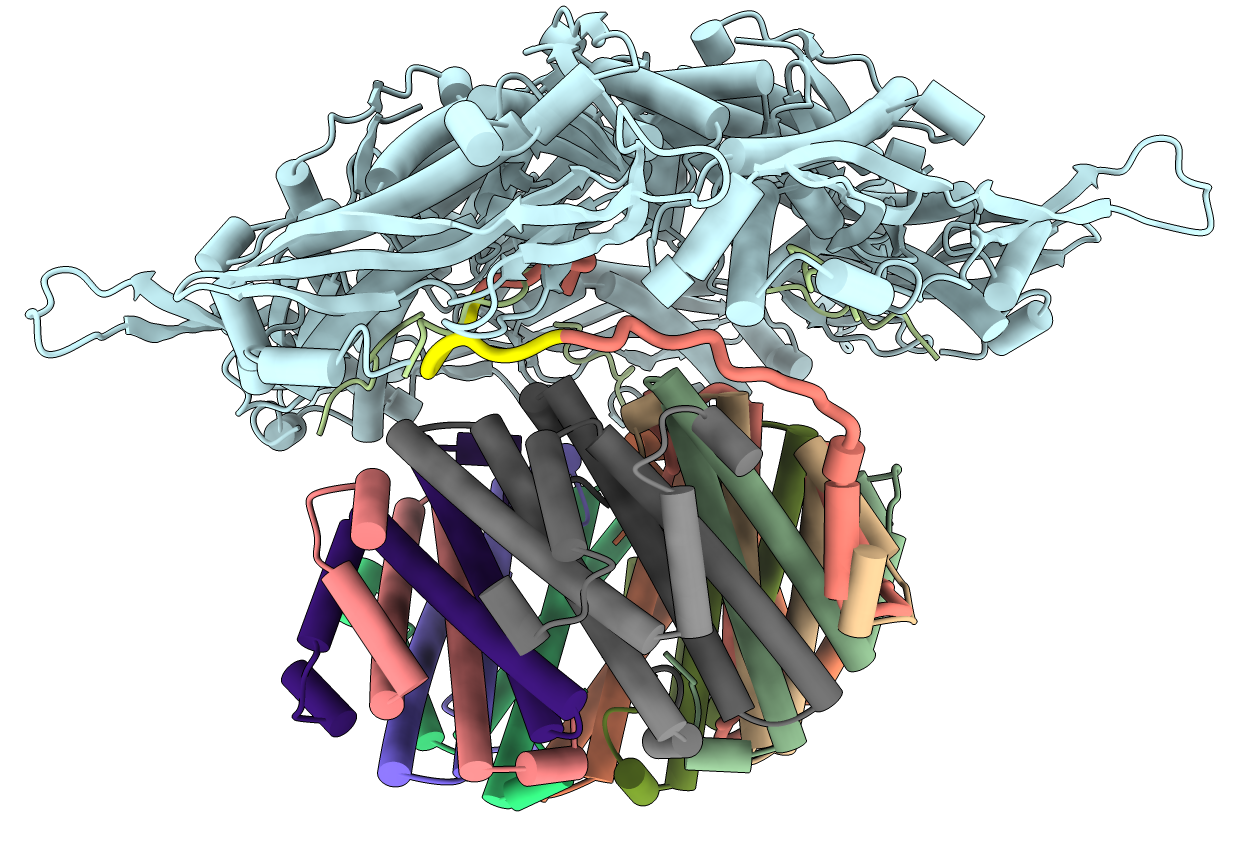
- Going to try to move localization signal and linker to show the connection
between the ferritin complex and the shell.
- Hide all atoms and show ribbon.
- Show atoms of AlphaFold model as ball and stick by selecting it with Model Panel selection button and using molecule display toolbar.
- Show sequence of alphafold model, menu Tools / Sequence.
- Drag select localization signal sequence 9-mer is several residues from end and has sequence (117)GSLGIGSLR(125).
- Use Tug mouse mode to drag residues of linker to overlay localization signal.
Can drag 3 or 4 residues along chain one at a time into position.
Full Encapsulin Ferritin Atomic Model
- Could do this for all 10 monomers in the ferritin complex and all 4 complexes to depict 40 linkers tethering to encapsulin shell.
- Would make full shell with 12 pentamers (60 monomers).
- 5 of 10 linkers exit the monomer pointing towards the middle of the shell and may
be hard to reach the shell.
Multi-Person VR
- Start a VR meeting by taking of headset, menu Tools / General / Meeting, meeting name "tg", person name Tom.
- Close the EMDB map. It is 500 Mbytes and would take a while to share over the network.
- Have Phil connect from East coast (I am in California).
- Explain Phil's head postage stamp shows where he is viewing and his cones show where he is pointing.
- Have Phil measure length of linker from protein to bound localization peptide using Tape Measure mouse mode.
- Show Phil and I both trying to move model making it vibrate -- there is no presenter, everyone has equal control.
- Multiple users can connect, we have tried only up to 4 person sessions.






