Pathway Extraction from Published Figures
Pathway information extracted from 25 years of pathway figures. Hanspers K, Riutta A, Summer-Kutmon M, Pico AR. Genome Biol. 2020 Nov 9;21(1):273. PMID: 33168034[back to paper list]
Identifying genes in published pathway figure images. Riutta A, Hanspers K, Pico AR. bioRxiv preprint 2018; doi: https://doi.org/10.1101/379446
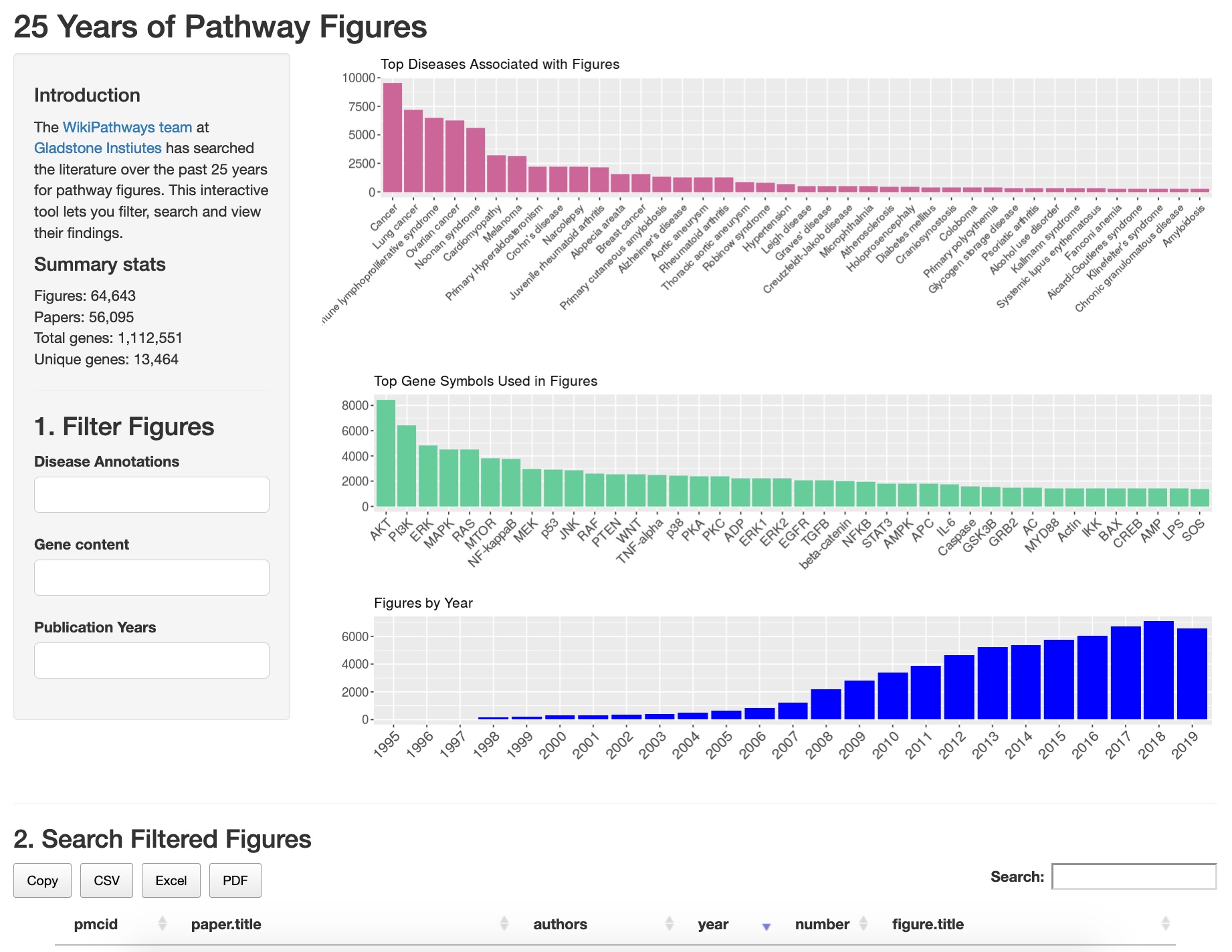
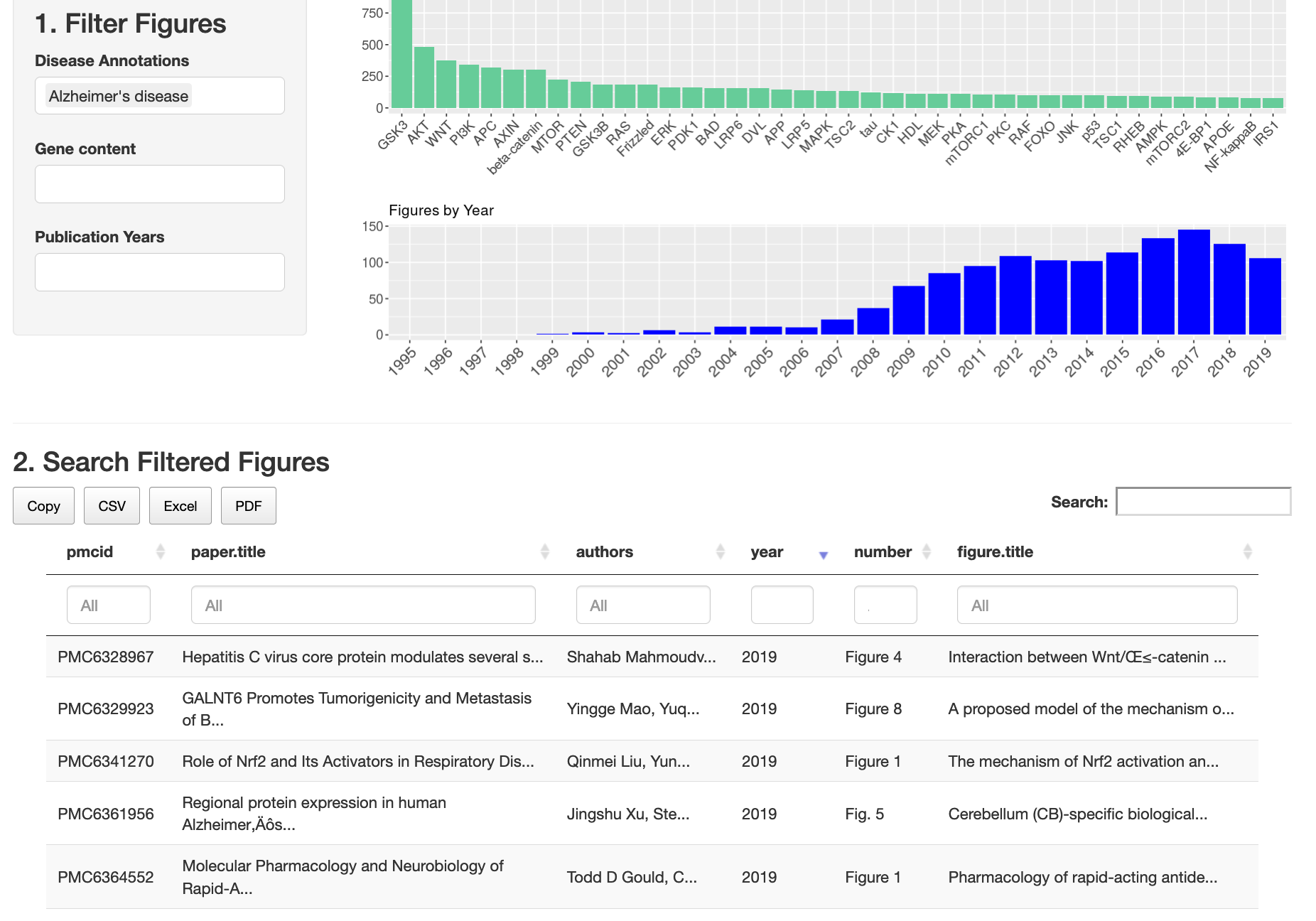
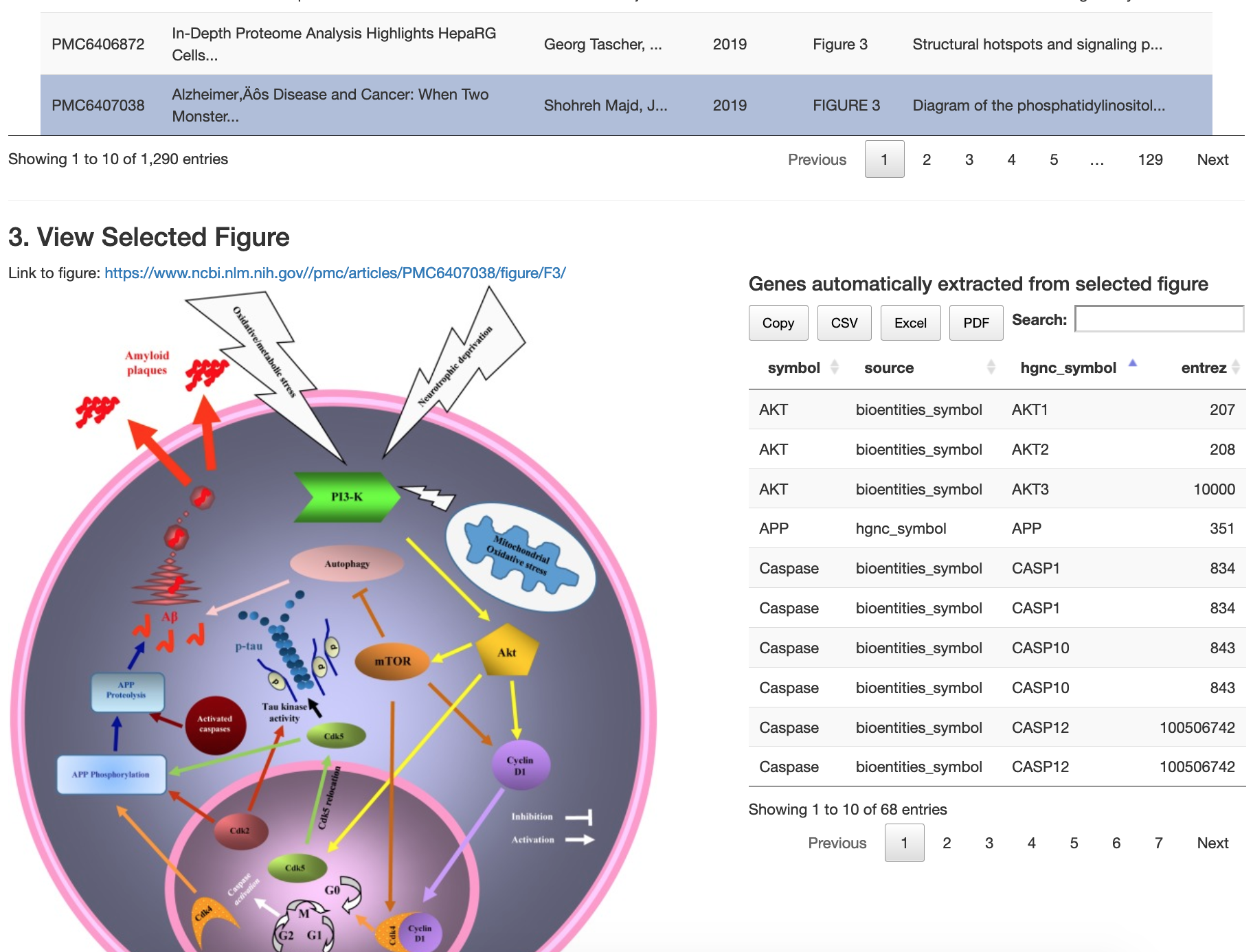
Web app for filtering, searching, and viewing the ~65K pathway figures:
https://gladstone-bioinformatics.shinyapps.io/shiny-25years
See also
https://gladstone-bioinformatics.shinyapps.io/shiny-covidpathways/
Bulk downloads of the pathway figures and OCR results are available on figshare.
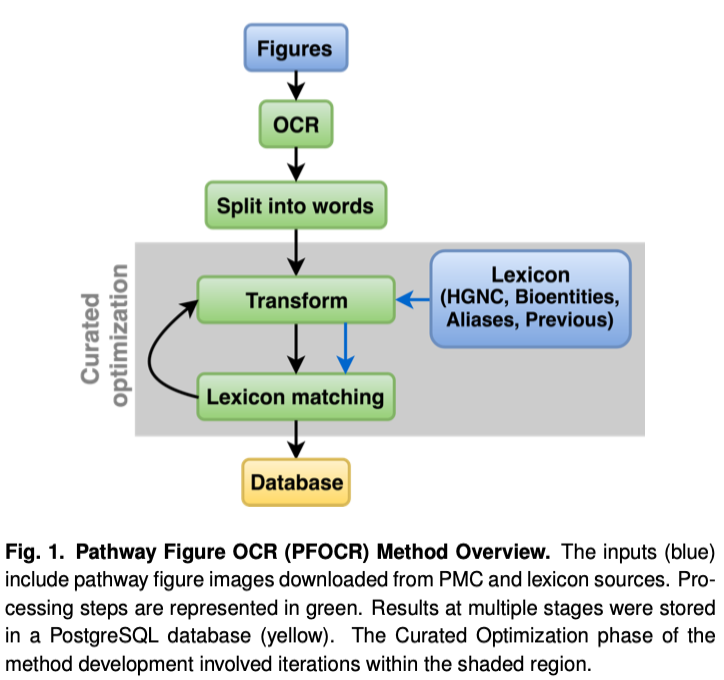
Overview
- Get images: PubMed Central (PMC) search
- Identify pathways: machine learning (ML) / computer vision
- Extract text: optical character recognition (OCR)
- Extract genes: rule-based named entity recognition (NER)
- Publish to [pathway] deposition targets
NLP – natural language processing
...switch over to Anders' slides from 3/5/2021...
(later should be available from https://wiki.library.ucsf.edu/display/NLPBiomed/NLP@UCSF+Meetups)
The lexicon includes four types of human gene symbols
mapped to NCBI Gene identifiers, from two sources:
Conflicts were resolved with priority order: HGNC symbol > bioentities > HGNC alias > HGNC previous. For example, if the same symbol from HGNC symbol and HGNC alias mapped to different NCBI Gene IDs, then only the HGNC symbol mapping was included in the lexicon. After curated optimization, the lexicon maps 58,242 unique symbols to 19,176 unique IDs.
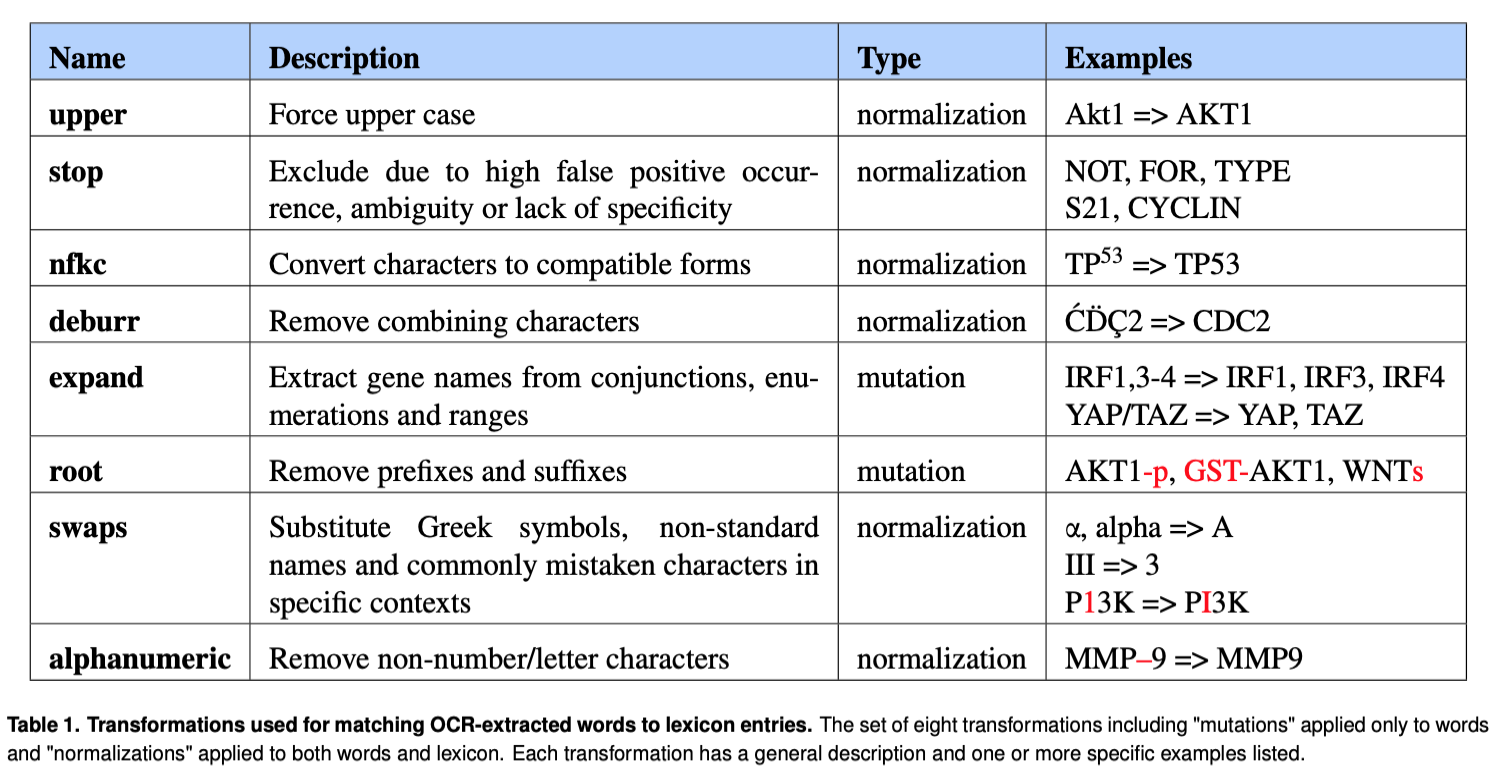
Transformations
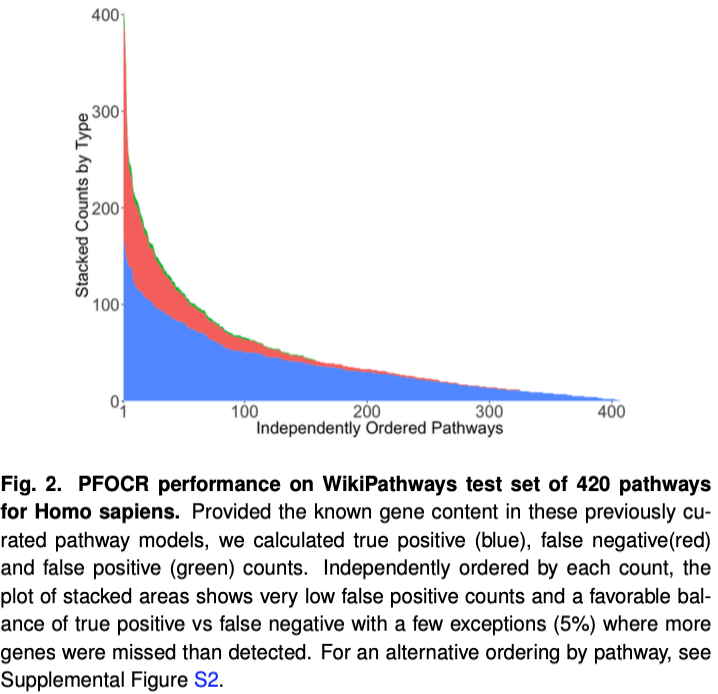
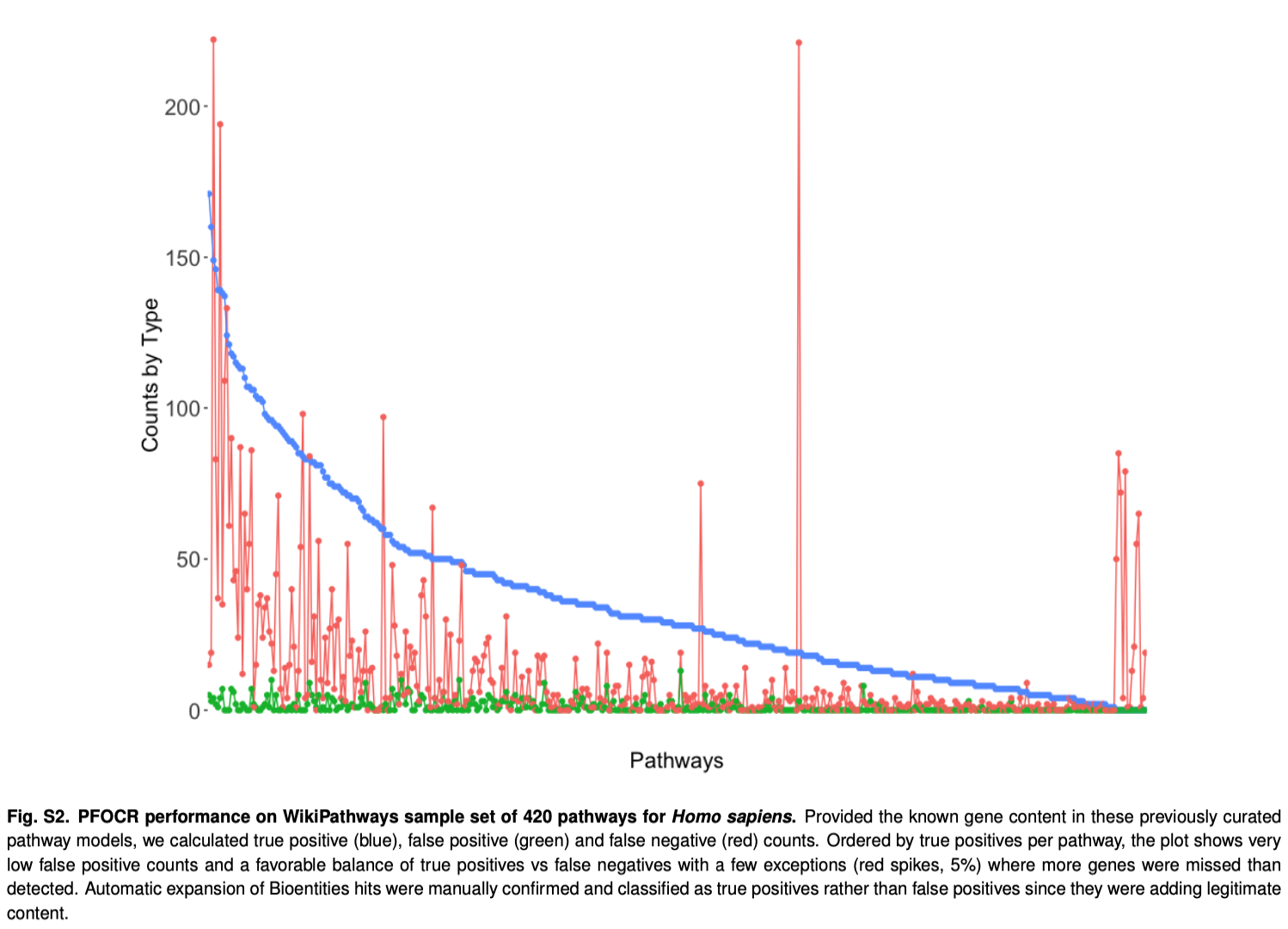
Validation on Curated Human Pathways
Statistics
- Get images: PubMed Central (PMC) search
- 253,081 figures from PMC image query using pathway-associated keywords (next slide) and pub date 1995-2019
- Identify pathways: machine learning (ML) / computer vision
- 64,643 pathway-likely figures (est. ~94% pathways) from ~56K papers in 3453 journals
- Extract text: optical character recognition (OCR)
- Extract genes: rule-based named entity recognition (NER)
- 58,962 figures with at least one human gene
- 1,112,551 instances of human genes, 13,464 unique human NCBI genes (average 18.9/figure)
- 28,836 figures with ≥ 7 human genes, nearly all of those figures associated with at least one GO biological process
- 20,227 figures associated with at least one disease ontology term, most commonly cancer
- Publish to [pathway] deposition targets
- currently working with NDEx to host the initial set of gene-annotated pathway figures for enrichment analysis
Pathway-associated keywords:
- pathway
- signaling
- regulatory
- disease
- drug
- metabolic
- biosynthetic
- synthesis
- cancer
- response
- cycle