UCSF ChimeraX
UCSF ChimeraX (or simply ChimeraX)
is the next-generation molecular visualization program from the
Resource for Biocomputing,
Visualization, and Informatics (RBVI),
following UCSF Chimera.
ChimeraX can be downloaded free of charge
for academic, government, nonprofit, and personal use.
Commercial users, please see
ChimeraX commercial licensing.
ChimeraX is developed with support from National Institutes of Health R01-GM129325.
 ChimeraX on Bluesky:
@chimerax.ucsf.edu
ChimeraX on Bluesky:
@chimerax.ucsf.edu
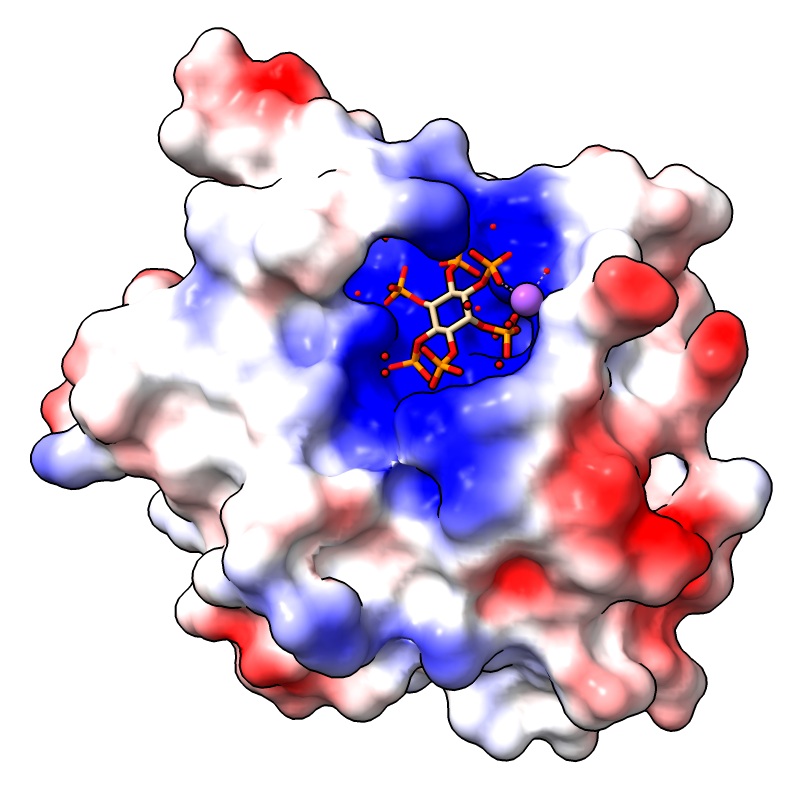
Coulombic electrostatic potential (ESP) can be calculated
and displayed with surface coloring using the command
coulombic
or the
Molecule Display
icon
 .
No separate calculation or input ESP file is required.
The image shows the first assembly defined for PDB
3eeb, the protease domain of a toxin from
Vibrio cholerae,
with the default Coulombic coloring: red-white-blue over the value range
–10 to 10. For image setup other than orientation, see the
command file coulombic.cxc.
.
No separate calculation or input ESP file is required.
The image shows the first assembly defined for PDB
3eeb, the protease domain of a toxin from
Vibrio cholerae,
with the default Coulombic coloring: red-white-blue over the value range
–10 to 10. For image setup other than orientation, see the
command file coulombic.cxc.
For how to add a color key and associated label, see the
Protein-Ligand Binding Sites tutorial.
More features...
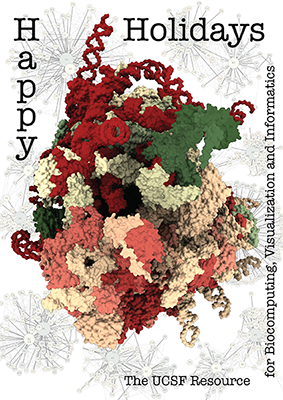
The architecture of the human ribosome has been determined at
near-atomic resolution by electron microscopy (Anger et al.,
Nature 497:80 (2013)).
The structure, comprising 82 proteins and five RNA molecules, is
shown with shadows cast from all directions to accentuate depth.
In the background are schematic representations of contacts
between the component molecules.
See the image setup script
card.cxc
using the
'Tis the Season color palette (credit to MrsP).
See also the RBVI
holiday card gallery.
More images...